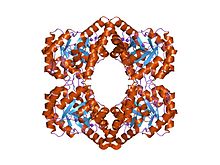
Lysine (symbol Lys or K) is an α-amino acid that is a precursor to many proteins. It contains an α-amino group (which is in the protonated −NH+
3 form under biological conditions), an α-carboxylic acid group (which is in the deprotonated −COO− form under biological conditions), and a side chain lysyl ((CH2)4NH2), classifying it as a basic, charged (at physiological pH), aliphatic amino acid. It is encoded by the codons AAA and AAG. Like almost all other amino acids, the α-carbon is chiral and lysine may refer to either enantiomer or a racemic mixture of both. For the purpose of this article, lysine will refer to the biologically active enantiomer L-lysine, where the α-carbon is in the S configuration.

Pyridoxal phosphate (PLP, pyridoxal 5'-phosphate, P5P), the active form of vitamin B6, is a coenzyme in a variety of enzymatic reactions. The International Union of Biochemistry and Molecular Biology has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. The versatility of PLP arises from its ability to covalently bind the substrate, and then to act as an electrophilic catalyst, thereby stabilizing different types of carbanionic reaction intermediates.

Oxaloacetic acid (also known as oxalacetic acid or OAA) is a crystalline organic compound with the chemical formula HO2CC(O)CH2CO2H. Oxaloacetic acid, in the form of its conjugate base oxaloacetate, is a metabolic intermediate in many processes that occur in animals. It takes part in gluconeogenesis, the urea cycle, the glyoxylate cycle, amino acid synthesis, fatty acid synthesis and the citric acid cycle.
In molecular biology, biosynthesis is a multi-step, enzyme-catalyzed process where substrates are converted into more complex products in living organisms. In biosynthesis, simple compounds are modified, converted into other compounds, or joined to form macromolecules. This process often consists of metabolic pathways. Some of these biosynthetic pathways are located within a single cellular organelle, while others involve enzymes that are located within multiple cellular organelles. Examples of these biosynthetic pathways include the production of lipid membrane components and nucleotides. Biosynthesis is usually synonymous with anabolism.

Amino acid synthesis is the set of biochemical processes by which the amino acids are produced. The substrates for these processes are various compounds in the organism's diet or growth media. Not all organisms are able to synthesize all amino acids. For example, humans can synthesize 11 of the 20 standard amino acids. These 11 are called the non-essential amino acids).
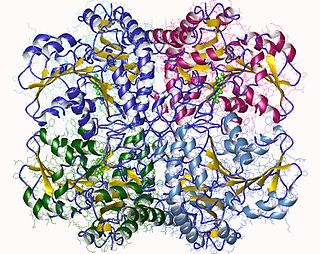
The enzyme cystathionine γ-lyase (EC 4.4.1.1, CTH or CSE; also cystathionase; systematic name L-cystathionine cysteine-lyase (deaminating; 2-oxobutanoate-forming)) breaks down cystathionine into cysteine, 2-oxobutanoate (α-ketobutyrate), and ammonia:
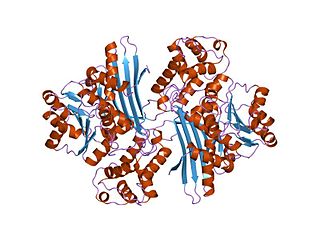
In molecular biology, the protein domain Saccharopine dehydrogenase (SDH), also named Saccharopine reductase, is an enzyme involved in the metabolism of the amino acid lysine, via an intermediate substance called saccharopine. The Saccharopine dehydrogenase enzyme can be classified under EC 1.5.1.7, EC 1.5.1.8, EC 1.5.1.9, and EC 1.5.1.10. It has an important function in lysine metabolism and catalyses a reaction in the alpha-Aminoadipic acid pathway. This pathway is unique to fungal organisms therefore, this molecule could be useful in the search for new antibiotics. This protein family also includes saccharopine dehydrogenase and homospermidine synthase. It is found in prokaryotes, eukaryotes and archaea.

The transsulfuration pathway is a metabolic pathway involving the interconversion of cysteine and homocysteine through the intermediate cystathionine. Two transsulfurylation pathways are known: the forward and the reverse.

In enzymology, an aspartate-semialdehyde dehydrogenase is an enzyme that is very important in the biosynthesis of amino acids in prokaryotes, fungi, and some higher plants. It forms an early branch point in the metabolic pathway forming lysine, methionine, leucine and isoleucine from aspartate. This pathway also produces diaminopimelate which plays an essential role in bacterial cell wall formation. There is particular interest in ASADH as disabling this enzyme proves fatal to the organism giving rise to the possibility of a new class of antibiotics, fungicides, and herbicides aimed at inhibiting it.
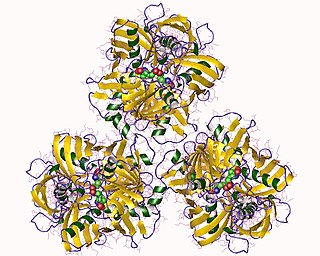
In enzymology, a diaminopimelate epimerase is an enzyme that catalyzes the chemical reaction

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction

The enzyme chorismate lyase catalyzes the first step in ubiquinone biosynthesis, the removal of pyruvate from chorismate, to yield 4-hydroxybenzoate in Escherichia coli and other Gram-negative bacteria. It belongs to the family of lyases, specifically the oxo-acid-lyases, which cleave carbon-carbon bonds. The systematic name of this enzyme class is chorismate pyruvate-lyase (4-hydroxybenzoate-forming). Other names in common use include CL, CPL, and UbiC.

The enzyme diaminopimelate decarboxylase (EC 4.1.1.20) catalyzes the cleavage of carbon-carbon bonds in meso 2,6 diaminoheptanedioate to produce CO2 and L-lysine, the essential amino acid. It employs the cofactor pyridoxal phosphate, also known as PLP, which participates in numerous enzymatic transamination, decarboxylation and deamination reactions.
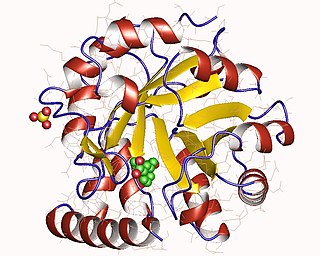
The enzyme indole-3-glycerol-phosphate synthase (IGPS) (EC 4.1.1.48) catalyzes the chemical reaction

Isocitrate lyase, or ICL, is an enzyme in the glyoxylate cycle that catalyzes the cleavage of isocitrate to succinate and glyoxylate. Together with malate synthase, it bypasses the two decarboxylation steps of the tricarboxylic acid cycle and is used by bacteria, fungi, and plants.
In enzymology, a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (EC 2.3.1.117) is an enzyme that catalyzes the chemical reaction

In enzymology, a malate synthase (EC 2.3.3.9) is an enzyme that catalyzes the chemical reaction
In enzymology, a N-acetylneuraminate synthase (EC 2.5.1.56) is an enzyme that catalyzes the chemical reaction
The bacterial cell wall provides strength and rigidity to counteract internal osmotic pressure, and protection against the environment. The peptidoglycan layer gives the cell wall its strength, and helps maintain the overall shape of the cell. The basic peptidoglycan structure of both Gram-positive and Gram-negative bacteria comprises a sheet of glycan chains connected by short cross-linking polypeptides. Biosynthesis of peptidoglycan is a multi-step process comprising three main stages:
- formation of UDP-N-acetylmuramic acid (UDPMurNAc) from N-acetylglucosamine (GlcNAc).
- addition of a short polypeptide chain to the UDPMurNAc.
- addition of a second GlcNAc to the disaccharide-pentapeptide building block and transport of this unit through the cytoplasmic membrane and incorporation into the growing peptidoglycan layer.

In enzymology, a 4-hydroxy-tetrahydrodipicolinate reductase (EC 1.17.1.8) is an enzyme that catalyzes the chemical reaction



















