
Pyruvate dehydrogenase complex (PDC) is a complex of three enzymes that converts pyruvate into acetyl-CoA by a process called pyruvate decarboxylation. Acetyl-CoA may then be used in the citric acid cycle to carry out cellular respiration, and this complex links the glycolysis metabolic pathway to the citric acid cycle. Pyruvate decarboxylation is also known as the "pyruvate dehydrogenase reaction" because it also involves the oxidation of pyruvate.
In molecular biology, biosynthesis is a multi-step, enzyme-catalyzed process where substrates are converted into more complex products in living organisms. In biosynthesis, simple compounds are modified, converted into other compounds, or joined to form macromolecules. This process often consists of metabolic pathways. Some of these biosynthetic pathways are located within a single cellular organelle, while others involve enzymes that are located within multiple cellular organelles. Examples of these biosynthetic pathways include the production of lipid membrane components and nucleotides. Biosynthesis is usually synonymous with anabolism.
The oxoglutarate dehydrogenase complex (OGDC) or α-ketoglutarate dehydrogenase complex is an enzyme complex, most commonly known for its role in the citric acid cycle.

Homoserine (also called isothreonine) is an α-amino acid with the chemical formula HO2CCH(NH2)CH2CH2OH. L-Homoserine is not one of the common amino acids encoded by DNA. It differs from the proteinogenic amino acid serine by insertion of an additional -CH2- unit into the backbone. Homoserine, or its lactone form, is the product of a cyanogen bromide cleavage of a peptide by degradation of methionine.
The branched-chain α-ketoacid dehydrogenase complex is a multi-subunit complex of enzymes that is found on the mitochondrial inner membrane. This enzyme complex catalyzes the oxidative decarboxylation of branched, short-chain alpha-ketoacids. BCKDC is a member of the mitochondrial α-ketoacid dehydrogenase complex family comprising pyruvate dehydrogenase and alpha-ketoglutarate dehydrogenase, key enzymes that function in the Krebs cycle.

Dihydrolipoyl transacetylase is an enzyme component of the multienzyme pyruvate dehydrogenase complex. The pyruvate dehydrogenase complex is responsible for the pyruvate decarboxylation step that links glycolysis to the citric acid cycle. This involves the transformation of pyruvate from glycolysis into acetyl-CoA which is then used in the citric acid cycle to carry out cellular respiration.
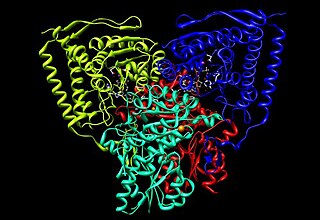
Pyruvate dehydrogenase is an enzyme that catalyzes the reaction of pyruvate and a lipoamide to give the acetylated dihydrolipoamide and carbon dioxide. The conversion requires the coenzyme thiamine pyrophosphate.
In enzymology, a succinylglutamate-semialdehyde dehydrogenase (EC 1.2.1.71) is an enzyme that catalyzes the chemical reaction

In enzymology, a 3-methyl-2-oxobutanoate dehydrogenase (EC 1.2.4.4) is an enzyme that catalyzes the chemical reaction

In enzymology, a saccharopine dehydrogenase (NAD+, L-lysine-forming) (EC 1.5.1.7) is an enzyme that catalyzes the chemical reaction
In enzymology, a N-succinylarginine dihydrolase (EC 3.5.3.23) is an enzyme that catalyzes the chemical reaction
In enzymology, a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase (EC 2.3.1.117) is an enzyme that catalyzes the chemical reaction
In enzymology, an arginine N-succinyltransferase (EC 2.3.1.109) is an enzyme that catalyzes the chemical reaction
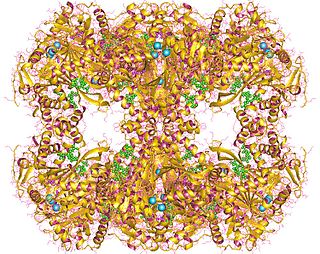

In enzymology, a dihydrolipoyllysine-residue (2-methylpropanoyl)transferase (EC 2.3.1.168) is an enzyme that catalyzes the chemical reaction

In enzymology, a homoserine O-succinyltransferase is an enzyme that catalyzes the chemical reaction
In enzymology, a lipoyl(octanoyl) transferase (EC 2.3.1.181) is an enzyme that catalyzes the chemical reaction

In enzymology, a succinylornithine transaminase (EC 2.6.1.81) is an enzyme that catalyzes the chemical reaction

Dihydrolipoyllysine-residue succinyltransferase component of 2-oxoglutarate dehydrogenase complex, mitochondrial is an enzyme that in humans is encoded by the DLST gene.

Alpha-ketoglutarate dehydrogenase also known as 2-oxoglutarate dehydrogenase E1 component, mitochondrial is an enzyme that in humans is encoded by the OGDH gene.
Lipoate–protein ligase (EC 2.7.7.63, LplA, lipoate protein ligase, lipoate–protein ligase A, LPL, LPL-B) is an enzyme with systematic name ATP:lipoate adenylyltransferase. This enzyme catalyses the following chemical reaction












