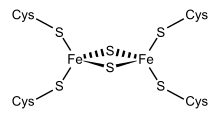
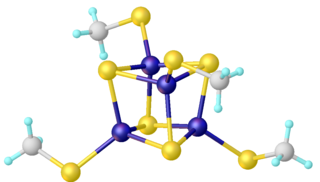
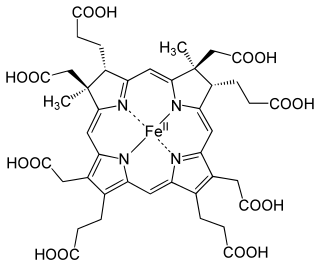
Bioenergetics of ferredoxins
Ferredoxins typically carry out a single electron transfer.
- Fd0
ox + e−Fd−
red
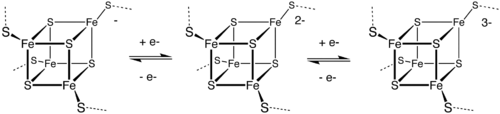
However a few bacterial ferredoxins (of the 2[4Fe4S] type) have two iron sulfur clusters and can carry out two electron transfer reactions. Depending on the sequence of the protein, the two transfers can have nearly identical reduction potentials or they may be significantly different. [4] [5]
- Fd0
ox + e−Fd−
red
- Fd−
red + e−Fd2−
red
Ferredoxins are one of the most reducing biological electron carriers. They typically have a mid point potential of -420 mV. [6] The reduction potential of a substance in the cell will differ from its midpoint potential depending on the concentrations of its reduced and oxidized forms. For a one electron reaction, the potential changes by around 60 mV for each power of ten change in the ratio of the concentration. For example, if the ferredoxin pool is around 95% reduced, the reduction potential will be around -500 mV. [7] In comparison, other biological reactions mostly have less reducing potentials: for example the primary biosynthetic reductant of the cell, NADPH has a cellular redox potential of -370 mV (E
0 = -320 mV).
Depending on the sequence of the supporting protein ferredoxins have reduction potential from around -500 mV [6] [8] to -340 mV. [9] A single cell can have multiple types of ferredoxins where each type is tuned to optimally carry out different reactions. [10]
Reduction of ferredoxin
The highly reducing ferredoxins are reduced either by using another strong reducing agent, or by using some source of energy to "boost" electrons from less reducing sources to the ferredoxin. [11]
Direct reduction
Reactions that reduce Fd include the oxidation of aldehydes to acids like the glyceraldehyde to glycerate reaction (-580 mV), the carbon monoxide dehydrogenase reaction (-520 mV), and the 2-oxoacid:Fd Oxidoreductase reactions (-500 mV) [12] [8] like the reaction carried out by pyruvate synthase. [7]
Membrane potential coupled reduction
Ferredoxin can also be reduced by using NADH (-320 mV) or H
2 (-414 mV), but these processes are coupled to the consumption of the membrane potential to power the "boosting" of electrons to the higher energy state. [6] The Rnf complex is a widespread membrane protein in bacteria that reversibly transfers electrons between NADH and ferredoxin while pumping Na+
or H+
ions across the cell membrane. The chemiosmotic potential of the membrane is consumed to power the unfavorable reduction of Fd
ox by NADH. This reaction is an essential source of Fd−
red in many autotrophic organisms. If the cell is growing on substrates that provide excess Fd−
red, the Rnf complex can transfer these electrons to NAD+
and store the resultant energy in the membrane potential. [13] The energy converting hydrogenases (Ech) are a family of enzymes that reversibly couple the transfer of electrons between Fd and H
2 while pumping H+
ions across the membrane to balance the energy difference. [14]
- Fd0
ox + NADH + Na+
outsideFd2−
red + NAD+
+ Na+
inside
- Fd0
ox + H
2 + H+
outsideFd2−
red + H+
+ H+
inside
Electron bifurcation
The unfavourable reduction of Fd from a less reducing electron donor can be coupled simultaneously with the favourable reduction of an oxidizing agent through an electron bifurcation reaction. [6] An example of the electron bifurcation reaction is the generation of Fd−
red for nitrogen fixation in certain aerobic diazotrophs. Typically, in oxidative phosphorylation the transfer of electrons from NADH to ubiquinone (Q) is coupled to charging the proton motive force. In Azotobacter the energy released by transferring one electron from NADH to Q is used to simultaneously boost the transfer of one electron from NADH to Fd. [15] [16]
Direct reduction of high potential ferredoxins
Some ferredoxins have a sufficiently high redox potential that they can be directly reduced by NADPH. One such ferredoxin is adrenoxin (-274 mV) which takes part in the biosynthesis of many mammalian steroids. [17] The ferredoxin Fd3 in the roots of plants that reduces nitrate and sulfite has a midpoint potential of -337 mV and is also reduced by NADPH. [10]