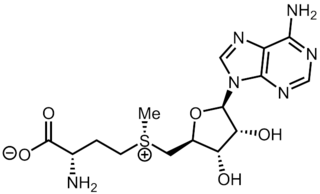
S-Adenosyl methionine (SAM), also known under the commercial names of SAMe, SAM-e, or AdoMet, is a common cosubstrate involved in methyl group transfers, transsulfuration, and aminopropylation. Although these anabolic reactions occur throughout the body, most SAM is produced and consumed in the liver. More than 40 methyl transfers from SAM are known, to various substrates such as nucleic acids, proteins, lipids and secondary metabolites. It is made from adenosine triphosphate (ATP) and methionine by methionine adenosyltransferase. SAM was first discovered by Giulio Cantoni in 1952.

Methyltransferases are a large group of enzymes that all methylate their substrates but can be split into several subclasses based on their structural features. The most common class of methyltransferases is class I, all of which contain a Rossmann fold for binding S-Adenosyl methionine (SAM). Class II methyltransferases contain a SET domain, which are exemplified by SET domain histone methyltransferases, and class III methyltransferases, which are membrane associated. Methyltransferases can also be grouped as different types utilizing different substrates in methyl transfer reactions. These types include protein methyltransferases, DNA/RNA methyltransferases, natural product methyltransferases, and non-SAM dependent methyltransferases. SAM is the classical methyl donor for methyltransferases, however, examples of other methyl donors are seen in nature. The general mechanism for methyl transfer is a SN2-like nucleophilic attack where the methionine sulfur serves as the leaving group and the methyl group attached to it acts as the electrophile that transfers the methyl group to the enzyme substrate. SAM is converted to S-Adenosyl homocysteine (SAH) during this process. The breaking of the SAM-methyl bond and the formation of the substrate-methyl bond happen nearly simultaneously. These enzymatic reactions are found in many pathways and are implicated in genetic diseases, cancer, and metabolic diseases. Another type of methyl transfer is the radical S-Adenosyl methionine (SAM) which is the methylation of unactivated carbon atoms in primary metabolites, proteins, lipids, and RNA.
Radical SAM is a designation for a superfamily of enzymes that use a [4Fe-4S]+ cluster to reductively cleave S-adenosyl-L-methionine (SAM) to generate a radical, usually a 5′-deoxyadenosyl radical (5'-dAdo), as a critical intermediate. These enzymes utilize this radical intermediate to perform diverse transformations, often to functionalize unactivated C-H bonds. Radical SAM enzymes are involved in cofactor biosynthesis, enzyme activation, peptide modification, post-transcriptional and post-translational modifications, metalloprotein cluster formation, tRNA modification, lipid metabolism, biosynthesis of antibiotics and natural products etc. The vast majority of known radical SAM enzymes belong to the radical SAM superfamily, and have a cysteine-rich motif that matches or resembles CxxxCxxC. rSAMs comprise the largest superfamily of metal-containing enzymes.
23S rRNA (uridine2552-2'-O)-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (uridine2552-2'-O-)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanine2445-N2)-methyltransferase (EC 2.1.1.173, ycbY (gene), rlmL (gene)) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanine2445-N2)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanine1835-N2)-methyltransferase (EC 2.1.1.174, ygjO (gene), rlmG (gene), ribosomal RNA large subunit methyltransferase G) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanine1835-N2)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (pseudouridine1915-N3)-methyltransferase (EC 2.1.1.177, YbeA, RlmH, pseudouridine methyltransferase, m3Psi methyltransferase, Psi1915-specific methyltransferase, rRNA large subunit methyltransferase H) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (pseudouridine1915-N3)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (adenine1618-N6)-methyltransferase (EC 2.1.1.181, rRNA large subunit methyltransferase F, YbiN protein, rlmF (gene), m6A1618 methyltransferase) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (adenine1618-N6)-methyltransferase. This enzyme catalyses the following chemical reaction
16S rRNA (adenine1518-N6/adenine1519-N6)-dimethyltransferase (EC 2.1.1.182, S-adenosylmethionine-6-N',N'-adenosyl (rRNA) dimethyltransferase, KsgA, ksgA methyltransferase) is an enzyme with systematic name S-adenosyl-L-methionine:16S rRNA (adenine1518-N6/adenine1519-N6)-dimethyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (adenine2085-N6)-dimethyltransferase (EC 2.1.1.184, ErmC' methyltransferase, ermC methylase, ermC 23S rRNA methyltransferase, rRNA:m6A methyltransferase ErmC', ErmC', rRNA methyltransferase ErmC' ) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (adenine2085-N6)-dimethyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanosine2251-2'-O)-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanosine2251-2'-O-)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (cytidine2498-2'-O)-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (cytidine2498-2'-O-)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanine745-N1)-methyltransferase (EC 2.1.1.187, Rlma(I), Rlma1, 23S rRNA m1G745 methyltransferase, YebH, RlmAI methyltransferase, ribosomal RNA(m1G)-methylase, rRNA(m1G)methylase, RrmA, 23S rRNA:m1G745 methyltransferase) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanine745-N1)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanine748-N1)-methyltransferase (EC 2.1.1.188, Rlma(II), Rlma2, 23S rRNA m1G748 methyltransferase, RlmaII, Rlma II, tylosin-resistance methyltransferase RlmA(II), TlrB, rRNA large subunit methyltransferase II) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanine748-N1)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (cytosine1962-C5)-methyltransferase (EC 2.1.1.191, RlmI, rRNA large subunit methyltransferase I, YccW) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (cytosine1962-C5)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (adenine2503-C2,C8)-dimethyltransferase (EC 2.1.1.194, Cfr, Cfr methyltransferase, Cfr rRNA methyltransferase) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (adenine2503-C2,C8)-dimethyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (adenine2503-C8)-methyltransferase (EC 2.1.1.224, Cfr (gene)) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (adenine2503-C8)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (cytidine1920-2'-O)-methyltransferase is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (cytidine1920-2'-O)-methyltransferase. This enzyme catalyses the following chemical reaction
16S rRNA (cytidine1409-2'-O)-methyltransferase (EC 2.1.1.227, TlyA) is an enzyme with systematic name S-adenosyl-L-methionine:16S rRNA (cytidine1409-2'-O)-methyltransferase. This enzyme catalyses the following chemical reaction
23S rRNA (guanine2069-N7)-methyltransferase (EC 2.1.1.264, rlmK (gene), 23S rRNA m7G2069 methyltransferase) is an enzyme with systematic name S-adenosyl-L-methionine:23S rRNA (guanine2069-N7)-methyltransferase. This enzyme catalyses the following chemical reaction


