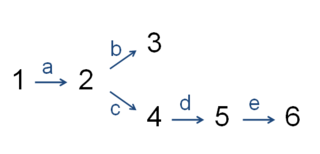The citric acid cycle —also known as the Krebs cycle, Szent-Györgyi-Krebs cycle or the TCA cycle (tricarboxylic acid cycle)—is a series of chemical reactions to release stored energy through the oxidation of acetyl-CoA derived from carbohydrates, fats, and proteins. The Krebs cycle is used by organisms that respire (as opposed to organisms that ferment) to generate energy, either by anaerobic respiration or aerobic respiration. In addition, the cycle provides precursors of certain amino acids, as well as the reducing agent NADH, that are used in numerous other reactions. Its central importance to many biochemical pathways suggests that it was one of the earliest components of metabolism. Even though it is branded as a 'cycle', it is not necessary for metabolites to follow only one specific route; at least three alternative segments of the citric acid cycle have been recognized.
The urea cycle (also known as the ornithine cycle) is a cycle of biochemical reactions that produces urea (NH2)2CO from ammonia (NH3). Animals that use this cycle, mainly amphibians and mammals, are called ureotelic.
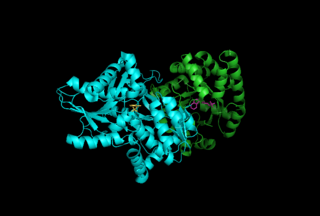
Tryptophan synthase or tryptophan synthetase is an enzyme that catalyses the final two steps in the biosynthesis of tryptophan. It is commonly found in Eubacteria, Archaebacteria, Protista, Fungi, and Plantae. However, it is absent from Animalia. It is typically found as an α2β2 tetramer. The α subunits catalyze the reversible formation of indole and glyceraldehyde-3-phosphate (G3P) from indole-3-glycerol phosphate (IGP). The β subunits catalyze the irreversible condensation of indole and serine to form tryptophan in a pyridoxal phosphate (PLP) dependent reaction. Each α active site is connected to a β active site by a 25 angstrom long hydrophobic channel contained within the enzyme. This facilitates the diffusion of indole formed at α active sites directly to β active sites in a process known as substrate channeling. The active sites of tryptophan synthase are allosterically coupled.
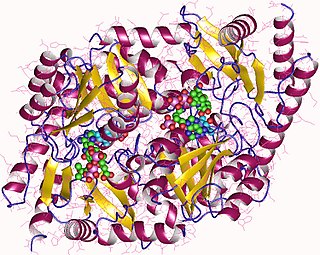
Aminolevulinic acid synthase (ALA synthase, ALAS, or delta-aminolevulinic acid synthase) is an enzyme (EC 2.3.1.37) that catalyzes the synthesis of δ-aminolevulinic acid (ALA) the first common precursor in the biosynthesis of all tetrapyrroles such as hemes, cobalamins and chlorophylls. The reaction is as follows:

Pyridoxal phosphate (PLP, pyridoxal 5'-phosphate, P5P), the active form of vitamin B6, is a coenzyme in a variety of enzymatic reactions. The International Union of Biochemistry and Molecular Biology has catalogued more than 140 PLP-dependent activities, corresponding to ~4% of all classified activities. The versatility of PLP arises from its ability to covalently bind the substrate, and then to act as an electrophilic catalyst, thereby stabilizing different types of carbanionic reaction intermediates.
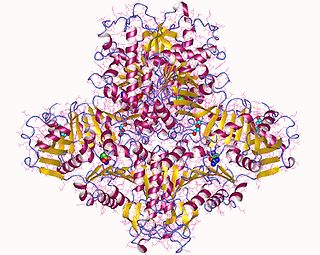
N-Acetylglutamate synthase (NAGS) is an enzyme that catalyses the production of N-acetylglutamate (NAG) from glutamate and acetyl-CoA.

Amino acid synthesis is the set of biochemical processes by which the amino acids are produced. The substrates for these processes are various compounds in the organism's diet or growth media. Not all organisms are able to synthesize all amino acids. For example, humans can synthesize 11 of the 20 standard amino acids. These 11 are called the non-essential amino acids).
Carbamoyl phosphate synthetase (glutamine-hydrolysing) is an enzyme that catalyzes the reactions that produce carbamoyl phosphate in the cytosol. Its systemic name is hydrogen-carbonate:L-glutamine amido-ligase .

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction

The enzyme methionine γ-lyase (EC 4.4.1.11, MGL) is in the γ-family of PLP-dependent enzymes. It degrades sulfur-containing amino acids to α-keto acids, ammonia, and thiols:

In enzymology, a D-alanine—D-alanine ligase is an enzyme that catalyzes the chemical reaction
Phosphoribosylamine—glycine ligase, also known as glycinamide ribonucleotide synthetase (GARS), (EC 6.3.4.13) is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-tripeptide—D-alanyl-D-alanine ligase is an enzyme that catalyzes the chemical reaction

The enzyme 3-dehydroquinate synthase catalyzes the chemical reaction

In enzymology, a serine C-palmitoyltransferase (EC 2.3.1.50) is an enzyme that catalyzes the chemical reaction:
In enzymology, a cysteine synthase is an enzyme that catalyzes the chemical reaction
In enzymology, an adenosylmethionine-8-amino-7-oxononanoate transaminase is an enzyme that catalyzes the chemical reaction
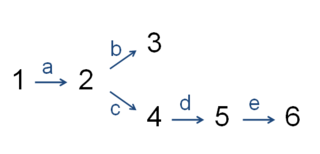
In enzymology, the committed step is an effectively irreversible enzymatic reaction that occurs at a branch point during the biosynthesis of some molecules. As the name implies, after this step, the molecules are "committed" to the pathway and will ultimately end up in the pathway's final product. The first committed step should not be confused with the rate-determining step, which is the slowest step in a reaction or pathway. However, it is sometimes the case that the first committed step is in fact the rate-determining step as well.
3-Deoxy-D-arabinoheptulosonate 7-phosphate (DAHP) synthase is the first enzyme in a series of metabolic reactions known as the shikimate pathway, which is responsible for the biosynthesis of the amino acids phenylalanine, tyrosine, and tryptophan. Since it is the first enzyme in the shikimate pathway, it controls the amount of carbon entering the pathway. Enzyme inhibition is the primary method of regulating the amount of carbon entering the pathway. Forms of this enzyme differ between organisms, but can be considered DAHP synthase based upon the reaction that is catalyzed by this enzyme.