
Oxidative phosphorylation or electron transport-linked phosphorylation or terminal oxidation is the metabolic pathway in which cells use enzymes to oxidize nutrients, thereby releasing chemical energy in order to produce adenosine triphosphate (ATP). In eukaryotes, this takes place inside mitochondria. Almost all aerobic organisms carry out oxidative phosphorylation. This pathway is so pervasive because it releases more energy than alternative fermentation processes such as anaerobic glycolysis.
A dehydrogenase is an enzyme belonging to the group of oxidoreductases that oxidizes a substrate by reducing an electron acceptor, usually NAD+/NADP+ or a flavin coenzyme such as FAD or FMN. Like all catalysts, they catalyze reverse as well as forward reactions, and in some cases this has physiological significance: for example, alcohol dehydrogenase catalyzes the oxidation of ethanol to acetaldehyde in animals, but in yeast it catalyzes the production of ethanol from acetaldehyde.

An electron transport chain (ETC) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. The electrons that are transferred from NADH and FADH2 to the ETC involves four multi-subunit large enzymes complexes and two mobile electron carriers. Many of the enzymes in the electron transport chain are embedded within the membrane.
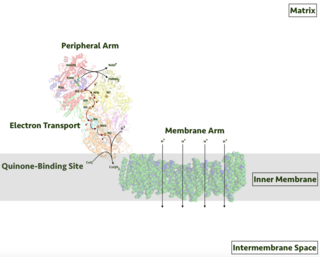
Respiratory complex I, EC 7.1.1.2 is the first large protein complex of the respiratory chains of many organisms from bacteria to humans. It catalyzes the transfer of electrons from NADH to coenzyme Q10 (CoQ10) and translocates protons across the inner mitochondrial membrane in eukaryotes or the plasma membrane of bacteria.
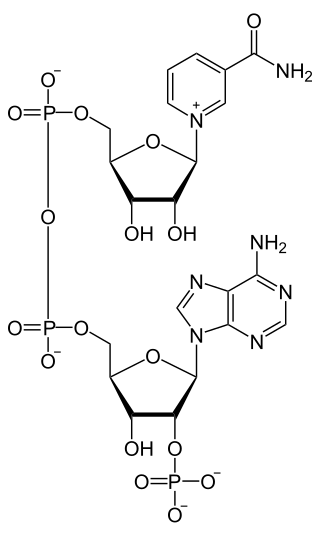
Nicotinamide adenine dinucleotide phosphate, abbreviated NADP+ or, in older notation, TPN (triphosphopyridine nucleotide), is a cofactor used in anabolic reactions, such as the Calvin cycle and lipid and nucleic acid syntheses, which require NADPH as a reducing agent ('hydrogen source'). NADPH is the reduced form of NADP+, the oxidized form. NADP+ is used by all forms of cellular life.

In biochemistry, flavin adenine dinucleotide (FAD) is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. A flavoprotein is a protein that contains a flavin group, which may be in the form of FAD or flavin mononucleotide (FMN). Many flavoproteins are known: components of the succinate dehydrogenase complex, α-ketoglutarate dehydrogenase, and a component of the pyruvate dehydrogenase complex.

Glutaryl-CoA dehydrogenase (GCDH) is an enzyme encoded by the GCDH gene on chromosome 19. The protein belongs to the acyl-CoA dehydrogenase family (ACD). It catalyzes the oxidative decarboxylation of glutaryl-CoA to crotonyl-CoA and carbon dioxide in the degradative pathway of L-lysine, L-hydroxylysine, and L-tryptophan metabolism. It uses electron transfer flavoprotein as its electron acceptor. The enzyme exists in the mitochondrial matrix as a homotetramer of 45-kD subunits. Mutations in this gene result in the metabolic disorder glutaric aciduria type 1, which is also known as glutaric acidemia type I. Alternative splicing of this gene results in multiple transcript variants.
Amine Dehydrogenase, also known as methylamine dehydrogenase (MADH), is a tryptophan tryptophylquinone-dependent (TTQ-dependent) enzyme that catalyzes the oxidative deamination of a primary amine to an aldehyde and ammonia. The reaction occurs as follows:
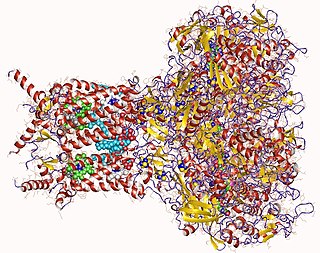
Formate dehydrogenases are a set of enzymes that catalyse the oxidation of formate to carbon dioxide, donating the electrons to a second substrate, such as NAD+ in formate:NAD+ oxidoreductase (EC 1.17.1.9) or to a cytochrome in formate:ferricytochrome-b1 oxidoreductase (EC 1.2.2.1). This family of enzymes has attracted attention as inspiration or guidance on methods for the carbon dioxide fixation, relevant to global warming.

Electron-transferring-flavoprotein dehydrogenase is an enzyme that transfers electrons from electron-transferring flavoprotein in the mitochondrial matrix, to the ubiquinone pool in the inner mitochondrial membrane. It is part of the electron transport chain. The enzyme is found in both prokaryotes and eukaryotes and contains a flavin and FE-S cluster. In humans, it is encoded by the ETFDH gene. Deficiency in ETF dehydrogenase causes the human genetic disease multiple acyl-CoA dehydrogenase deficiency.
In enzymology, a quinoprotein glucose dehydrogenase is an enzyme that catalyzes the chemical reaction
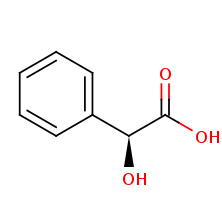
In enzymology, (S)-mandelate dehydrogenase (MDH), is an enzyme that catalyzes the chemical reaction.
In enzymology, carbon monoxide dehydrogenase (CODH) (EC 1.2.7.4) is an enzyme that catalyzes the chemical reaction

Azurin is a small, periplasmic, bacterial blue copper protein found in Pseudomonas, Bordetella, or Alcaligenes bacteria. Azurin moderates single-electron transfer between enzymes associated with the cytochrome chain by undergoing oxidation-reduction between Cu(I) and Cu(II). Each monomer of an azurin tetramer has a molecular weight of approximately 14kDa, contains a single copper atom, is intensively blue, and has a fluorescence emission band centered at 308 nm.
Methanol dehydrogenase (cytochrome c) (EC 1.1.2.7, methanol dehydrogenase, MDH) is an enzyme with systematic name methanol:cytochrome c oxidoreductase. This enzyme catalyses the following chemical reaction
Alcohol dehydrogenase (azurin) (EC 1.1.9.1, type II quinoprotein alcohol dehydrogenase, quinohaemoprotein ethanol dehydrogenase, QHEDH, ADHIIB) is an enzyme with systematic name alcohol:azurin oxidoreductase. This enzyme catalyses the following chemical reaction
Soluble quinoprotein glucose dehydrogenase is an enzyme with systematic name D-glucose:acceptor oxidoreductase. This enzyme catalyses the following chemical reaction
Long-chain acyl-CoA dehydrogenase is an enzyme with systematic name long-chain acyl-CoA:electron-transfer flavoprotein 2,3-oxidoreductase. This enzyme catalyses the following chemical reaction
Methylamine dehydrogenase (amicyanin) (EC 1.4.9.1, amine dehydrogenase, primary-amine dehydrogenase) is an enzyme with systematic name methylamine:amicyanin oxidoreductase (deaminating). This enzyme catalyses the following chemical reaction:

NADH:ubiquinone reductase (non-electrogenic) (EC 1.6.5.9, NDH-2, ubiquinone reductase, coenzyme Q reductase, dihydronicotinamide adenine dinucleotide-coenzyme Q reductase, DPNH-coenzyme Q reductase, DPNH-ubiquinone reductase, NADH-coenzyme Q oxidoreductase, NADH-coenzyme Q reductase, NADH-CoQ oxidoreductase, NADH-CoQ reductase) is an enzyme with systematic name NADH:ubiquinone oxidoreductase. This enzyme catalyses the following chemical reaction:











