
In biochemistry, a ribonucleotide is a nucleotide containing ribose as its pentose component. It is considered a molecular precursor of nucleic acids. Nucleotides are the basic building blocks of DNA and RNA. Ribonucleotides themselves are basic monomeric building blocks for RNA. Deoxyribonucleotides, formed by reducing ribonucleotides with the enzyme ribonucleotide reductase (RNR), are essential building blocks for DNA. There are several differences between DNA deoxyribonucleotides and RNA ribonucleotides. Successive nucleotides are linked together via phosphodiester bonds.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

The TIM barrel, also known as an alpha/beta barrel, is a conserved protein fold consisting of eight alpha helices (α-helices) and eight parallel beta strands (β-strands) that alternate along the peptide backbone. The structure is named after triose-phosphate isomerase, a conserved metabolic enzyme. TIM barrels are ubiquitous, with approximately 10% of all enzymes adopting this fold. Further, five of seven enzyme commission (EC) enzyme classes include TIM barrel proteins. The TIM barrel fold is evolutionarily ancient, with many of its members possessing little similarity today, instead falling within the twilight zone of sequence similarity.
Phosphoribosylformylglycinamidine cyclo-ligase is the fifth enzyme in the de novo synthesis of purine nucleotides. It catalyzes the reaction to form 5-aminoimidazole ribotide (AIR) from formylglycinamidine-ribonucleotide FGAM. This reaction closes the ring and produces a 5-membered imidazole ring of the purine nucleus (AIR):
Purine metabolism refers to the metabolic pathways to synthesize and break down purines that are present in many organisms.

Guanosine monophosphate synthetase, also known as GMPS is an enzyme that converts xanthosine monophosphate to guanosine monophosphate.

In molecular biology, adenylosuccinate synthase is an enzyme that plays an important role in purine biosynthesis, by catalysing the guanosine triphosphate (GTP)-dependent conversion of inosine monophosphate (IMP) and aspartic acid to guanosine diphosphate (GDP), phosphate and N(6)-(1,2-dicarboxyethyl)-AMP. Adenylosuccinate synthetase has been characterised from various sources ranging from Escherichia coli to vertebrate tissues. In vertebrates, two isozymes are present: one involved in purine biosynthesis and the other in the purine nucleotide cycle.

Amidophosphoribosyltransferase (ATase), also known as glutamine phosphoribosylpyrophosphate amidotransferase (GPAT), is an enzyme responsible for catalyzing the conversion of 5-phosphoribosyl-1-pyrophosphate (PRPP) into 5-phosphoribosyl-1-amine (PRA), using the amine group from a glutamine side-chain. This is the committing step in de novo purine synthesis. In humans it is encoded by the PPAT gene. ATase is a member of the purine/pyrimidine phosphoribosyltransferase family.

Bifunctional purine biosynthesis protein PURH is a protein that in humans is encoded by the ATIC gene.
In enzymology, a 5-(carboxyamino)imidazole ribonucleotide mutase is an enzyme that catalyzes the chemical reaction

In enzymology, a phosphoribosylanthranilate isomerase (PRAI) is an enzyme that catalyzes the third step of the synthesis of the amino acid tryptophan.
In enzymology, a 5-(carboxyamino)imidazole ribonucleotide synthase (EC 6.3.4.18) is an enzyme that catalyzes the chemical reaction
Phosphoribosylamine—glycine ligase, also known as glycinamide ribonucleotide synthetase (GARS), (EC 6.3.4.13) is an enzyme that catalyzes the chemical reaction
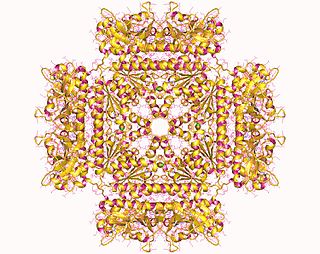

In molecular biology, the protein domain SAICAR synthase is an enzyme which catalyses a reaction to create SAICAR. In enzymology, this enzyme is also known as phosphoribosylaminoimidazolesuccinocarboxamide synthase. It is an enzyme that catalyzes the chemical reaction

In enzymology, a phosphoribosylformylglycinamidine synthase (EC 6.3.5.3) is an enzyme that catalyzes the chemical reaction

In enzymology, a phosphoribosyl-AMP cyclohydrolase (EC 3.5.4.19) is an enzyme that catalyzes the chemical reaction

In enzymology, an anthranilate phosphoribosyltransferase is an enzyme that catalyzes the chemical reaction

In enzymology, an ATP phosphoribosyltransferase is an enzyme that catalyzes the chemical reaction
Phosphoribosylglycinamide formyltransferase (EC 2.1.2.2), also known as glycinamide ribonucleotide transformylase (GAR Tfase), is an enzyme with systematic name 10-formyltetrahydrofolate:5'-phosphoribosylglycinamide N-formyltransferase. This enzyme catalyses the following chemical reaction

In molecular biology, the YjeF N terminal is a protein domain found in the N-terminal of the protein, EDC3. The YjeF N-terminal domains occur either as single proteins or fusions with other domains and are commonly associated with enzymes. They help assemble the processing body (P-body) in preparation for mRNAdecay. Structural homology indicated it may have some similarity to the enzyme family, hydrolase.

















