
Biological databases are libraries of biological sciences, collected from scientific experiments, published literature, high-throughput experiment technology, and computational analysis. They contain information from research areas including genomics, proteomics, metabolomics, microarray gene expression, and phylogenetics. Information contained in biological databases includes gene function, structure, localization, clinical effects of mutations as well as similarities of biological sequences and structures.
BRENDA is an information system representing one of the most comprehensive enzyme repositories. It is an electronic resource that comprises molecular and biochemical information on enzymes that have been classified by the IUBMB. Every classified enzyme is characterized with respect to its catalyzed biochemical reaction. Kinetic properties of the corresponding reactants are described in detail. BRENDA contains enzyme-specific data manually extracted from primary scientific literature and additional data derived from automatic information retrieval methods such as text mining. It provides a web-based user interface that allows a convenient and sophisticated access to the data.
In academia, computational immunology is a field of science that encompasses high-throughput genomic and bioinformatics approaches to immunology. The field's main aim is to convert immunological data into computational problems, solve these problems using mathematical and computational approaches and then convert these results into immunologically meaningful interpretations.

Amos Bairoch is a Swiss bioinformatician and Professor of Bioinformatics at the Department of Human Protein Sciences of the University of Geneva where he leads the CALIPHO group at the Swiss Institute of Bioinformatics (SIB) combining bioinformatics, curation, and experimental efforts to functionally characterize human proteins.
The Saccharomyces Genome Database (SGD) is a scientific database of the molecular biology and genetics of the yeast Saccharomyces cerevisiae, which is commonly known as baker's or budding yeast.
The MetaCyc database is one of the largest metabolic pathways and enzymes databases currently available. The data in the database is manually curated from the scientific literature, and covers all domains of life. MetaCyc has extensive information about chemical compounds, reactions, metabolic pathways and enzymes. The data have been curated from more than 58,000 publications.

The Integrated Microbial Genomes system is a genome browsing and annotation platform developed by the U.S. Department of Energy (DOE)-Joint Genome Institute. IMG contains all the draft and complete microbial genomes sequenced by the DOE-JGI integrated with other publicly available genomes. IMG provides users a set of tools for comparative analysis of microbial genomes along three dimensions: genes, genomes and functions. Users can select and transfer them in the comparative analysis carts based upon a variety of criteria. IMG also includes a genome annotation pipeline that integrates information from several tools, including KEGG, Pfam, InterPro, and the Gene Ontology, among others. Users can also type or upload their own gene annotations and the IMG system will allow them to generate Genbank or EMBL format files containing these annotations.
Mouse Genome Informatics (MGI) is a free, online database and bioinformatics resource hosted by The Jackson Laboratory, with funding by the National Human Genome Research Institute (NHGRI), the National Cancer Institute (NCI), and the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD). MGI provides access to data on the genetics, genomics and biology of the laboratory mouse to facilitate the study of human health and disease. The database integrates multiple projects, with the two largest contributions coming from the Mouse Genome Database and Mouse Gene Expression Database (GXD). As of 2018, MGI contains data curated from over 230,000 publications.
PDBsum is a database that provides an overview of the contents of each 3D macromolecular structure deposited in the Protein Data Bank. The original version of the database was developed around 1995 by Roman Laskowski and collaborators at University College London. As of 2014, PDBsum is maintained by Laskowski and collaborators in the laboratory of Janet Thornton at the European Bioinformatics Institute (EBI).
CharProtDB is a curated database of biochemically characterized proteins.

The Human Metabolome Database (HMDB) is a comprehensive, high-quality, freely accessible, online database of small molecule metabolites found in the human body. Created by the Human Metabolome Project funded by Genome Canada. One of the first dedicated metabolomics databases, the HMDB facilitates human metabolomics research, including the identification and characterization of human metabolites using NMR spectroscopy, GC-MS spectrometry and LC/MS spectrometry. To aid in this discovery process, the HMDB contains three kinds of data: 1) chemical data, 2) clinical data, and 3) molecular biology/biochemistry data. The chemical data includes 41,514 metabolite structures with detailed descriptions along with nearly 10,000 NMR, GC-MS and LC/MS spectra.
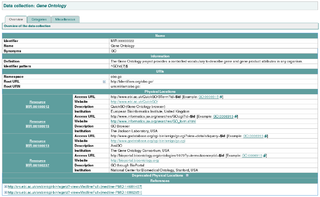
The MIRIAM Registry, a by-product of the MIRIAM Guidelines, is a database of namespaces and associated information that is used in the creation of uniform resource identifiers. It contains the set of community-approved namespaces for databases and resources serving, primarily, the biological sciences domain. These shared namespaces, when combined with 'data collection' identifiers, can be used to create globally unique identifiers for knowledge held in data repositories. For more information on the use of URIs to annotate models, see the specification of SBML Level 2 Version 2.

Experimental factor ontology, also known as EFO, is an open-access ontology of experimental variables particularly those used in molecular biology. The ontology covers variables which include aspects of disease, anatomy, cell type, cell lines, chemical compounds and assay information. EFO is developed and maintained at the EMBL-EBI as a cross-cutting resource for the purposes of curation, querying and data integration in resources such as Ensembl, ChEMBL and Expression Atlas.
In bioinformatics, a Gene Disease Database is a systematized collection of data, typically structured to model aspects of reality, in a way to comprehend the underlying mechanisms of complex diseases, by understanding multiple composite interactions between phenotype-genotype relationships and gene-disease mechanisms. Gene Disease Databases integrate human gene-disease associations from various expert curated databases and text mining derived associations including Mendelian, complex and environmental diseases.
The Expression Atlas is a database maintained by the European Bioinformatics Institute that provides information on gene expression patterns from RNA-Seq and Microarray studies, and protein expression from Proteomics studies. The Expression Atlas allows searches by gene, splice variant, protein attribute, disease, treatment or organism part. Individual genes or gene sets can be searched for. All datasets in Expression Atlas have its metadata manually curated and its data analysed through standardised analysis pipelines. There are two components to the Expression Atlas, the Baseline Atlas and the Differential Atlas:

BacDive is a bacterial metadatabase that provides strain-linked information about bacterial and archaeal biodiversity.
Model organism databases (MODs) are biological databases, or knowledgebases, dedicated to the provision of in-depth biological data for intensively studied model organisms. MODs allow researchers to easily find background information on large sets of genes, plan experiments efficiently, combine their data with existing knowledge, and construct novel hypotheses. They allow users to analyse results and interpret datasets, and the data they generate are increasingly used to describe less well studied species. Where possible, MODs share common approaches to collect and represent biological information. For example, all MODs use the Gene Ontology (GO) to describe functions, processes and cellular locations of specific gene products. Projects also exist to enable software sharing for curation, visualization and querying between different MODs. Organismal diversity and varying user requirements however mean that MODs are often required to customize capture, display, and provision of data.

Biocuration is the field of life sciences dedicated to organizing biomedical data, information and knowledge into structured formats, such as spreadsheets, tables and knowledge graphs. The biocuration of biomedical knowledge is made possible by the cooperative work of biocurators, software developers and bioinformaticians and is at the base of the work of biological databases.








