See also
Y-DNA Q-M242 Subclades
Y-DNA backbone tree
| |||
Footnotes
| |||
| Haplogroup Q-P89.1 | |
|---|---|
| Possible time of origin | Insufficient data [1] |
| Possible place of origin | Asia or Beringia |
| Ancestor | Q-MEH2 |
| Defining mutations | P89.1 |
Haplogroup Q-P89.1 is a subclade of Y-DNA Haplogroup Q-MEH2. [1] Haplogroup Q-P89.1 is defined by the presence of the P89.1 Single Nucleotide Polymorphism (SNP). In 2010, Q-P89.1 was reclassified as "private" and removed from the haplotree. [2]
Q-P89.1 has descendants in the Northwest Territory of modern Canada. It was in pre-Columbian American populations that it was discovered. [1] [3]
Q-P89.1 is present in pre-Columbian populations in the Canadian Northwest. [1]
| Population | Paper | N | Percentage | SNP Tested | |
|---|---|---|---|---|---|
| Gwich’in | Dulik 2012 | 0/33 | ~0.00% | P89.1 | |
| Tłįchǫ | Dulik 2012 | 1/37 | ~2.70% | P89.1 | |
| Inuvialuit | Dulik 2012 | 0/56 | ~0.00% | P89.1 | |
| Inupiat | Dulik 2012 | 0/5 | ~0.00% | P89.1 |
Because samples from Asia have only sporadically been tested for this lineage, its frequency there is uncertain.
Q-P89.1 is currently defined by only the P89.1 SNP.

Haplogroup J-M304, also known as J, is a human Y-chromosome DNA haplogroup. It is believed to have evolved in Western Asia. The clade spread from there during the Neolithic, primarily into North Africa, the Horn of Africa, the Socotra Archipelago, the Caucasus, Europe, Anatolia, Central Asia, South Asia, and Southeast Asia.

Haplogroup F, also known as F-M89 and previously as Haplogroup FT, is a very common Y-chromosome haplogroup. The clade and its subclades constitute over 90% of paternal lineages outside of Africa.
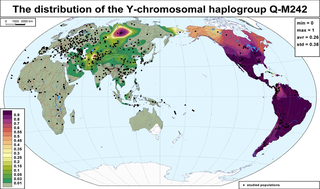
Haplogroup Q or Q-M242 is a Y-chromosome DNA haplogroup. It has one primary subclade, Haplogroup Q1 (L232/S432), which includes numerous subclades that have been sampled and identified in males among modern populations.
Haplogroup Q-M3 (Y-DNA) is a Y-chromosome DNA haplogroup. Haplogroup Q-M3 is a subclade of Haplogroup Q-L54. Haplogroup Q-M3 was previously known as Haplogroup Q3; currently Q-M3 is Q1b1a1a below Q1b-M346.

In human genetics, a human Y-chromosome DNA haplogroup is a haplogroup defined by mutations in the non-recombining portions of DNA from the male-specific Y chromosome. Many people within a haplogroup share similar numbers of short tandem repeats (STRs) and types of mutations called single-nucleotide polymorphisms (SNPs).

Haplogroup NO1, also known as NO-M214, is a human Y-chromosome DNA haplogroup. NO1 is the sole confirmed subclade of Haplogroup K- M2313, which is the sole subclade of Haplogroup K2a (K-M2308). NO is the dominant Y-DNA haplogroup in most parts of eastern and northern Eurasia, including East Asia, Siberia and northern Fennoscandia.

Haplogroup CT is a human Y chromosome haplogroup. CT has two basal branches, CF and DE. DE is divided into a predominantly Asia-distributed haplogroup D-CTS3946 and a predominantly Africa-distributed haplogroup E-M96, while CF is divided into an East Asian, Native American, and Oceanian haplogroup C-M130 and haplogroup F-M89, which dominates most non-African populations.
In human population genetics, Y-Chromosome haplogroups define the major lineages of direct paternal (male) lines back to a shared common ancestor in Africa. Men in the same haplogroup share a set of differences, or markers, on their Y-Chromosome, which distinguish them from men in other haplogroups. These UEPs, or markers used to define haplogroups, are SNP mutations. Y-Chromosome Haplogroups all form "family trees" or "phylogenies", with both branches or sub-clades diverging from a common haplogroup ancestor, and also with all haplogroups themselves linked into one family tree which traces back ultimately to the most recent shared male line ancestor of all men alive today, called in popular science Y Chromosome Adam.
Haplogroup Q-M346 is a subclade of Y-DNA Haplogroup Q. Haplogroup Q-M346 is defined by the presence of the M346 Single Nucleotide Polymorphism (SNP).
Haplogroup Q-L54 is a subclade of Y-DNA haplogroup Q-L53. Q1a3a-L54 is defined by the presence of the L54 Single Nucleotide Polymorphism (SNP).
Haplogroup Q-L275 or Haplogroup Q2 is a human Y-chromosome DNA haplogroup believed to have originated in Eurasia. Haplogroup Q-L275 is defined by the presence of the L275 single-nucleotide polymorphism (SNP). Haplogroup Q-L275 can be identified through genealogical DNA testing.
Haplogroup Q-NWT01 is a subclade of Y-DNA Haplogroup Q-MEH2. Haplogroup Q-NWT01 is defined by the presence of the NWT01 Single Nucleotide Polymorphism (SNP).
Haplogroup Q-M25, also known as Q1a1b is a subclade or branch of human Y-DNA haplogroup Q-F1096 (Q1a1), which is, in turn, a subclade of Q-MEH2 (Q1a). In human genetics, each Y-DNA haplogroup constitutes a biological paternal lineages back to a shared common male ancestor.
Haplogroup Q-M120, also known as Q1a1a1, is a Y-DNA haplogroup. It is the only primary branch of haplogroup Q1a1a (F746/NWT01). The lineage is most common amongst modern populations in eastern Eurasia.
Q-L53 is a subclade of haplogroup Q-M346. Q-L53 is defined by the presence of the L53 Single Nucleotide Polymorphism (SNP).
Haplogroup Q-Z780 is a subclade of the Y-DNA Haplogroup Q-L54. Q-Z780 is defined by the presence of the Z780 Single Nucleotide Polymorphism (SNP).
Haplogroup Q-M323 is a subclade of Y-DNA Haplogroup Q-M346. Haplogroup Q-M323 is defined by the presence of the M323 Single Nucleotide Polymorphism (SNP).
HaplogroupGHIJK, defined by the SNPs M3658, F1329, PF2622, and YSC0001299, is a common Y-chromosome haplogroup. This macrohaplogroup and its subclades contain the vast majority of the world's existing male population.

Haplogroup R-M269 is the sub-clade of human Y-chromosome haplogroup R1b that is defined by the SNP marker M269. According to ISOGG 2020 it is phylogenetically classified as R1b1a1b. It underwent intensive research and was previously classified as R1b1a2, R1b1c, R1b1b2 and R1b1a1a2.
Haplogroup Q-L804 (Y-DNA) is a Y-chromosome DNA haplogroup. Haplogroup Q-L804 is a subclade of Haplogroup Q-L54. Currently Q-L804 is Q1b1a1b below Q1b-M346.