Related Research Articles

A monoclonal antibody is an antibody produced from a cell lineage made by cloning a unique white blood cell. All subsequent antibodies derived this way trace back to a unique parent cell.

A polyhistidine-tag, best known by the trademarked name His-tag, is an amino acid motif in proteins that typically consists of at least six histidine (His) residues, often at the N- or C-terminus of the protein. It is also known as a hexa histidine-tag, 6xHis-tag, or His6 tag. The tag was invented by Roche, although the use of histidines and its vectors are distributed by Qiagen. Various purification kits for histidine-tagged proteins are commercially available from multiple companies.
Protein purification is a series of processes intended to isolate one or a few proteins from a complex mixture, usually cells, tissues or whole organisms. Protein purification is vital for the specification of the function, structure and interactions of the protein of interest. The purification process may separate the protein and non-protein parts of the mixture, and finally separate the desired protein from all other proteins. Ideally, to study a protein of interest, it must be separated from other components of the cell so that contaminants will not interfere in the examination of the protein of interest's structure and function. Separation of one protein from all others is typically the most laborious aspect of protein purification. Separation steps usually exploit differences in protein size, physico-chemical properties, binding affinity and biological activity. The pure result may be termed protein isolate.
In biochemistry, biotinylation is the process of covalently attaching biotin to a protein, nucleic acid or other molecule. Biotinylation is rapid, specific and is unlikely to disturb the natural function of the molecule due to the small size of biotin. Biotin binds to streptavidin and avidin with an extremely high affinity, fast on-rate, and high specificity, and these interactions are exploited in many areas of biotechnology to isolate biotinylated molecules of interest. Biotin-binding to streptavidin and avidin is resistant to extremes of heat, pH and proteolysis, making capture of biotinylated molecules possible in a wide variety of environments. Also, multiple biotin molecules can be conjugated to a protein of interest, which allows binding of multiple streptavidin, avidin or neutravidin protein molecules and increases the sensitivity of detection of the protein of interest. There is a large number of biotinylation reagents available that exploit the wide range of possible labelling methods. Due to the strong affinity between biotin and streptavidin, the purification of biotinylated proteins has been a widely used approach to identify protein-protein interactions and post-translational events such as ubiquitylation in molecular biology.
Affinity chromatography is a method of separating a biomolecule from a mixture, based on a highly specific macromolecular binding interaction between the biomolecule and another substance. The specific type of binding interaction depends on the biomolecule of interest; antigen and antibody, enzyme and substrate, receptor and ligand, or protein and nucleic acid binding interactions are frequently exploited for isolation of various biomolecules. Affinity chromatography is useful for its high selectivity and resolution of separation, compared to other chromatographic methods.
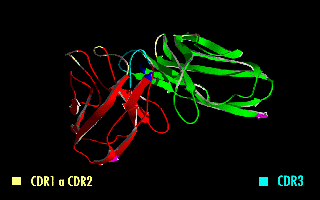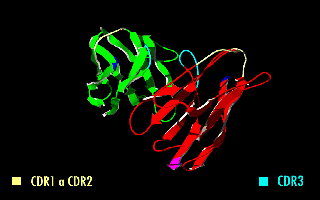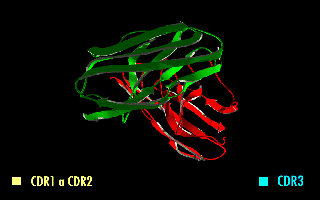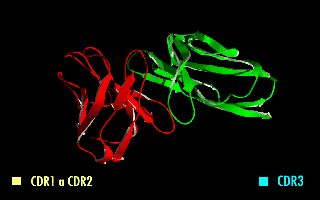
A single-chain variable fragment (scFv) is not actually a fragment of an antibody, but instead is a fusion protein of the variable regions of the heavy (VH) and light chains (VL) of immunoglobulins, connected with a short linker peptide of ten to about 25 amino acids. The linker is usually rich in glycine for flexibility, as well as serine or threonine for solubility, and can either connect the N-terminus of the VH with the C-terminus of the VL, or vice versa. This protein retains the specificity of the original immunoglobulin, despite removal of the constant regions and the introduction of the linker. The image to the right shows how this modification usually leaves the specificity unaltered.

Streptavidin is a 52 kDa protein (tetramer) purified from the bacterium Streptomyces avidinii. Streptavidin homo-tetramers have an extraordinarily high affinity for biotin. With a dissociation constant (Kd) on the order of ≈10−14 mol/L, the binding of biotin to streptavidin is one of the strongest non-covalent interactions known in nature. Streptavidin is used extensively in molecular biology and bionanotechnology due to the streptavidin-biotin complex's resistance to organic solvents, denaturants, detergents, proteolytic enzymes, and extremes of temperature and pH.
Protein tags are peptide sequences genetically grafted onto a recombinant protein. Tags are attached to proteins for various purposes. They can be added to either end of the target protein, so they are either C-terminus or N-terminus specific or are both C-terminus and N-terminus specific. Some tags are also inserted at sites within the protein of interest; they are known as internal tags.
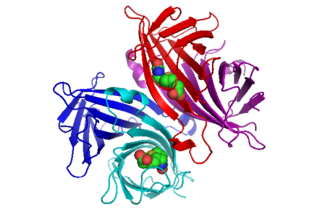
Avidin is a tetrameric biotin-binding protein produced in the oviducts of birds, reptiles and amphibians and deposited in the whites of their eggs. Dimeric members of the avidin family are also found in some bacteria. In chicken egg white, avidin makes up approximately 0.05% of total protein (approximately 1800 μg per egg). The tetrameric protein contains four identical subunits (homotetramer), each of which can bind to biotin (Vitamin B7, vitamin H) with a high degree of affinity and specificity. The dissociation constant of the avidin-biotin complex is measured to be KD ≈ 10−15 M, making it one of the strongest known non-covalent bonds.

Protein A is a 42 kDa surface protein originally found in the cell wall of the bacteria Staphylococcus aureus. It is encoded by the spa gene and its regulation is controlled by DNA topology, cellular osmolarity, and a two-component system called ArlS-ArlR. It has found use in biochemical research because of its ability to bind immunoglobulins. It is composed of five homologous Ig-binding domains that fold into a three-helix bundle. Each domain is able to bind proteins from many mammalian species, most notably IgGs. It binds the heavy chain within the Fc region of most immunoglobulins and also within the Fab region in the case of the human VH3 family. Through these interactions in serum, where IgG molecules are bound in the wrong orientation, the bacteria disrupts opsonization and phagocytosis.
Multicolumn countercurrent solvent gradient purification (MCSGP) is a form of chromatography that is used to separate or purify biomolecules from complex mixtures. It was developed at the Swiss Federal Institute of Technology Zürich by Aumann and Morbidelli. The process consists of two to six chromatographic columns which are connected to one another in such a way that as the mixture moves through the columns the compound is purified into several fractions.
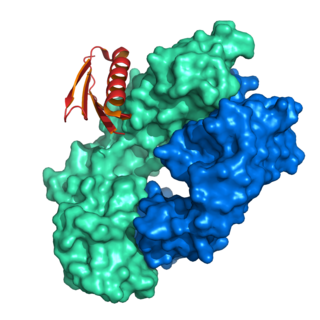
Protein L was first isolated from the surface of bacterial species Peptostreptococcus magnus and was found to bind immunoglobulins through L chain interaction, from which the name was suggested. It consists of 719 amino acid residues. The molecular weight of protein L purified from the cell walls of Peptostreptoccus magnus was first estimated as 95kD by SDS-PAGE in the presence of reducing agent 2-mercaptoethanol, while the molecular weight was determined to 76kD by gel chromatography in the presence of 6 M guanidine HCl. Protein L does not contain any interchain disulfide loops, nor does it consist of disulfide-linked subunits. It is an acidic molecule with a pI of 4.0. Unlike protein A and protein G, which bind to the Fc region of immunoglobulins (antibodies), protein L binds antibodies through light chain interactions. Since no part of the heavy chain is involved in the binding interaction, Protein L binds a wider range of antibody classes than protein A or G. Protein L binds to representatives of all antibody classes, including IgG, IgM, IgA, IgE and IgD. Single chain variable fragments (scFv) and Fab fragments also bind to protein L.
Protein A/G is a recombinant fusion protein that combines IgG binding domains of both protein A and protein G. Protein A/G contains four Fc binding domains from protein A and two from protein G, yielding a final mass of 50,460 daltons. The binding of protein A/G is less pH-dependent than protein A, but otherwise has the additive properties of protein A and G.

Meir Wilchek is an Israeli biochemist. He is a professor at the Weizmann Institute of Science.
The Strep-tag system is a method which allows the purification and detection of proteins by affinity chromatography. The Strep-tag II is a synthetic peptide consisting of eight amino acids (Trp-Ser-His-Pro-Gln-Phe-Glu-Lys). This peptide sequence exhibits intrinsic affinity towards Strep-Tactin, a specifically engineered streptavidin, and can be N- or C- terminally fused to recombinant proteins. By exploiting the highly specific interaction, Strep-tagged proteins can be isolated in one step from crude cell lysates. Because the Strep-tag elutes under gentle, physiological conditions, it is especially suited for the generation of functional proteins.
Affibody molecules are small, robust proteins engineered to bind to a large number of target proteins or peptides with high affinity, imitating monoclonal antibodies, and are therefore a member of the family of antibody mimetics. Affibody molecules are used in biochemical research and are being developed as potential new biopharmaceutical drugs. These molecules can be used for molecular recognition in diagnostic and therapeutic applications.
Antibody mimetics are organic compounds that, like antibodies, can specifically bind antigens, but that are not structurally related to antibodies. They are usually artificial peptides or proteins with a molar mass of about 3 to 20 kDa.

Affitins are artificial proteins with the ability to selectively bind antigens. They are structurally derived from the DNA binding protein Sac7d, found in Sulfolobus acidocaldarius, a microorganism belonging to the archaeal domain. By randomizing the amino acids on the binding surface of Sac7d and subjecting the resulting protein library to rounds of ribosome display, the affinity can be directed towards various targets, such as peptides, proteins, viruses, and bacteria.
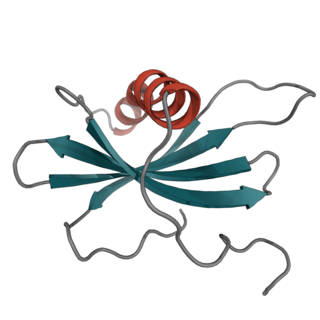
Affimer molecules are small proteins that bind to target proteins with affinity in the nanomolar range. These engineered non-antibody binding proteins are designed to mimic the molecular recognition characteristics of monoclonal antibodies in different applications. These affinity reagents have been optimized to increase their stability, make them tolerant to a range of temperatures and pH, reduce their size, and to increase their expression in E.coli and mammalian cells.
Sophia Hober is a Swedish researcher in biotechnology and professor at The Royal Institute of Technology (KTH) in Stockholm. Sophia Hober got her Master of Science in chemical engineering at KTH in 1989 and defended her doctorate in biochemistry in 1996. Since 2007, Hober is a professor of molecular biotechnology at KTH. During 2011–2015, Professor Hober served as dean at KTH and was part of the management team. Sophia Hober was elected member of the Royal Swedish Academy of Engineering Sciences in 2012.
References
- 1 2 3 4 5 6 Suderman RJ, Rice DA, Gibson SD, Strick EJ, Chao DM (Apr 17, 2017). "Development of polyol-responsive antibody mimetics for single-step protein purification". Protein Expression and Purification. 134: 114–124. doi: 10.1016/j.pep.2017.04.008 . PMID 28428153.
- 1 2 Burgess RR, Watson JD (June 2017). "Gentle antibody-mimetic affinity chromatography with polyol-responsive nanoCLAMPs". Protein Expression and Purification. 134: 154–155. doi:10.1016/j.pep.2017.06.002. PMID 28633910.
- 1 2 Thompson NE, Aronson DB, Burgess RR (1990). "Purification of eukaryotic RNA polymerase II by immunoaffinity chromatography. Elution of active enzyme with protein stabilizing agents from a polyol-responsive monoclonal antibody". J Biol Chem. 265 (12): 7069–7077. doi: 10.1016/S0021-9258(19)39260-9 . PMID 2324114.
- ↑ Thompson NE, Foley KM, Stalder ES, Burgess, RR (2009). "Chapter 28 Identification, Production, and Use of Polyol-Responsive Monoclonal Antibodies for Immunoaffinity Chromatography". Guide to Protein Purification, 2nd Edition. Methods Enzymol. Vol. 463. pp. 475–494. doi:10.1016/S0076-6879(09)63028-7. ISBN 978-0-12-374536-1. PMID 19892188.
- ↑ Thompson NE, Hager DA, Burgess RR (Aug 4, 1992). "Isolation and characterization of a polyol-responsive monoclonal antibody useful for gentle purification of Escherichia coli RNA polymerase". Biochemistry. 31 (30): 7003–7008. doi:10.1021/bi00145a019. PMID 1637835.
- ↑ Thompson NE, Jensen DB, Lamberski JA, Burgess RR (2006). "Purification of protein complexes by immunoaffinity chromatography: application to transcription machinery". Genet Eng (N Y). Genetic Engineering: Principles and Methods. 27: 81–100. doi:10.1007/0-387-25856-6_6. ISBN 0-387-25855-8. PMID 16382873.
