| Nuclear Pore | |
|---|---|
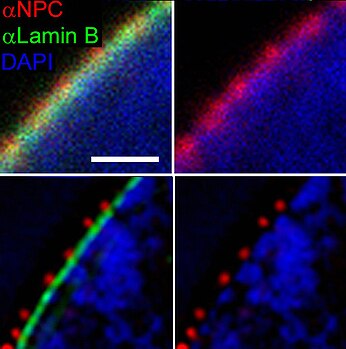
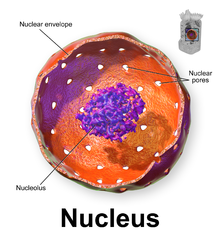
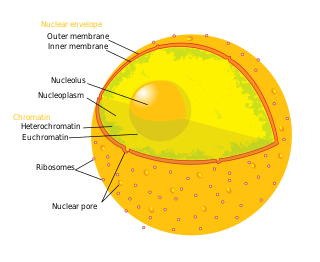
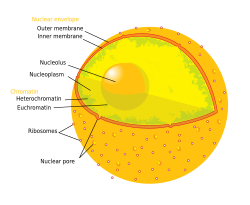 Diagram of the human cell nucleus with nuclear pores. | |
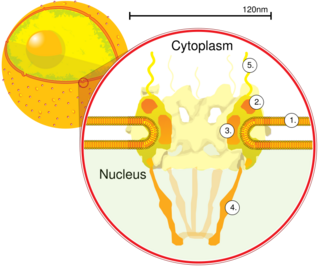
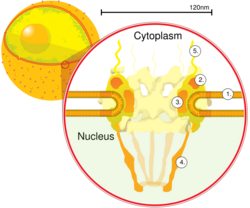 Schematic diagram of a nuclear pore complex within the nuclear envelope (1) with the outer ring (2), spokes (3), basket (4), and filaments (5). | |
| Details | |
| Identifiers | |
| Latin | porus nuclearis |
| MeSH | D022022 |
| TH | H1.00.01.2.01005 |
| FMA | 63148 |
| Anatomical terminology | |
A nuclear pore is a channel as part of the nuclear pore complex (NPC), a large protein complex found in the nuclear envelope of eukaryotic cells. The nuclear envelope (NE) surrounds the cell nucleus containing DNA and facilitates the selective membrane transport of various molecules.
Contents
- Nuclear pore complex
- Nuclear transport
- Import of proteins
- Export of proteins
- Export of RNA
- Assembly of the NPC
- Theories of assembly
- Disassembly
- Preservation of integrity
- References
- External links
The nuclear pore complex consists predominantly of proteins known as nucleoporins (Nups). Each human NPC is composed of about 1,000 individual protein molecules, from an evolutionarily conserved set of 35 distinct nucleoporins. [1] In 2022 around 90% of the structure of the human NPC was elucidated in an open and a closed conformation, and published in a special issue of Science, featured on the cover. [2] [3] [4] [5] In 2024 the structure of the nuclear basket was solved, finalising the structure of NPC. [6]
About half of the nucleoporins encompass solenoid protein domains, such as alpha solenoids or beta-propeller folds, and occasionally both as separate structural domains. Conversely, the remaining nucleoporins exhibit characteristics of "natively unfolded" or intrinsically disordered proteins, characterized by high flexibility and a lack of ordered tertiary structure. These disordered proteins, referred to as FG nucleoporins (FG-Nups), contain multiple phenylalanine–glycine repeats (FG repeats) in their amino acid sequences. [7] FG-Nups is one of three main types of nucleoporins found in the NPC. The other two are the transmembrane Nups and the scaffold Nups. The transmembrane Nups are made up of transmembrane α-helices and play a vital part in anchoring the NPC to the nuclear envelope. The scaffold Nups are made up of α-solenoid and β-propeller folds, and create the structural framework of NPCs. [8]
The principal function of nuclear pore complexes is to facilitate selective membrane transport of various molecules across the nuclear envelope. This includes the transportation of RNA and ribosomal proteins from the nucleus to the cytoplasm, as well as proteins (such as DNA polymerase and lamins), carbohydrates, signaling molecules, and lipids moving into the nucleus. Notably, the nuclear pore complex (NPC) can actively mediate up to 1000 translocations per complex per second. While smaller molecules can passively diffuse through the pores, larger molecules are often identified by specific signal sequences and are facilitated by nucleoporins to traverse the nuclear envelope.
Evolutionary conserved features in sequences that code for nucleoporins regulate molecular transport through the nuclear pore. [9] [10] Nucleoporin-mediated transport does not entail direct energy expenditure but instead relies on concentration gradients associated with the RAN cycle (Ras-related nuclear protein cycle).
The count of nuclear pore complexes varies across cell types and different stages of the cell's life cycle, with approximately 1,000 NPCs typically found in vertebrate cells. [11] The human nuclear pore complex (hNPC) is a substantial structure, with a molecular weight of 120 megadaltons (MDa). [12] Each NPC comprises eight protein subunits encircling the actual pore, forming the outer ring. Additionally, these subunits project a spoke-shaped protein over the pore channel. The central region of the pore may exhibit a plug-like structure; however, its precise nature remains unknown, and it is yet undetermined whether it represents an actual plug or merely cargo transiently caught in transit.