
MicroRNA (miRNA) are small, single-stranded, non-coding RNA molecules containing 21 to 23 nucleotides. Found in plants, animals and some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in mRNA molecules, then silence said mRNA molecules by one or more of the following processes:
- Cleavage of the mRNA strand into two pieces,
- Destabilization of the mRNA by shortening its poly(A) tail, or
- Reducing translation of the mRNA into proteins.

Sterol regulatory element-binding proteins (SREBPs) are transcription factors that bind to the sterol regulatory element DNA sequence TCACNCCAC. Mammalian SREBPs are encoded by the genes SREBF1 and SREBF2. SREBPs belong to the basic-helix-loop-helix leucine zipper class of transcription factors. Unactivated SREBPs are attached to the nuclear envelope and endoplasmic reticulum membranes. In cells with low levels of sterols, SREBPs are cleaved to a water-soluble N-terminal domain that is translocated to the nucleus. These activated SREBPs then bind to specific sterol regulatory element DNA sequences, thus upregulating the synthesis of enzymes involved in sterol biosynthesis. Sterols in turn inhibit the cleavage of SREBPs and therefore synthesis of additional sterols is reduced through a negative feed back loop.
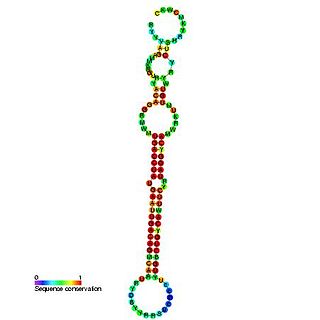
The miR-192 microRNA precursor, is a short non-coding RNA gene involved in gene regulation. miR-192 and miR-215 have now been predicted or experimentally confirmed in mouse and human.

The miR-1 microRNA precursor is a small micro RNA that regulates its target protein's expression in the cell. microRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give products at ~22 nucleotides. In this case the mature sequence comes from the 3' arm of the precursor. The mature products are thought to have regulatory roles through complementarity to mRNA. In humans there are two distinct microRNAs that share an identical mature sequence, and these are called miR-1-1 and miR-1-2.
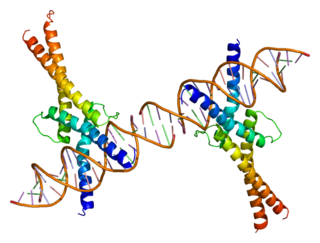
Sterol regulatory element-binding transcription factor 1 (SREBF1) also known as sterol regulatory element-binding protein 1 (SREBP-1) is a protein that in humans is encoded by the SREBF1 gene.
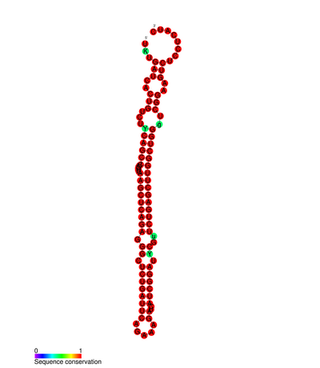
mir-127 microRNA is a short non-coding RNA molecule with interesting overlapping gene structure. miR-127 functions to regulate the expression levels of genes involved in lung development, placental formation and apoptosis. Aberrant expression of miR-127 has been linked to different cancers.

In molecular biology, mir-145 microRNA is a short RNA molecule that in humans is encoded by the MIR145 gene. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
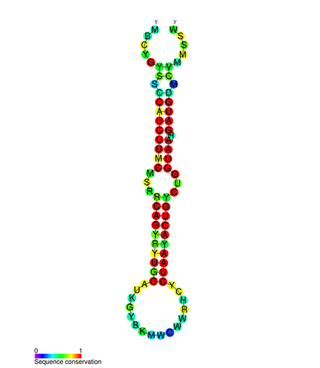
In molecular biology, the miR-200 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by binding and cleaving mRNAs or inhibiting translation. The miR-200 family contains miR-200a, miR-200b, miR-200c, miR-141, and miR-429. There is growing evidence to suggest that miR-200 microRNAs are involved in cancer metastasis.

In molecular biology MicroRNA-223 (miR-223) is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. miR-223 is a hematopoietic specific microRNA with crucial functions in myeloid lineage development. It plays an essential role in promoting granulocytic differentiation while also being associated with the suppression of erythrocytic differentiation. miR-223 is commonly repressed in hepatocellular carcinoma and leukemia. Higher expression levels of miRNA-223 are associated with extranodal marginal-zone lymphoma of mucosa-associated lymphoid tissue of the stomach and recurrent ovarian cancer. In some cancers the microRNA-223 down-regulation is correlated with higher tumor burden, disease aggressiveness, and poor prognostic factors. MicroRNA-223 is also associated with rheumatoid arthritis, sepsis, type 2 diabetes, and hepatic ischemia.
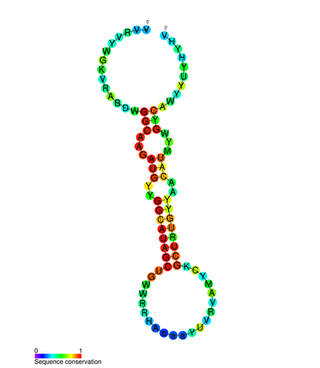
miR-31 has been characterised as a tumour suppressor miRNA, with its levels varying in breast cancer cells according to the metastatic state of the tumour. From its typical abundance in healthy tissue is a moderate decrease in non-metastatic breast cancer cell lines, and levels are almost completely absent in mouse and human metastatic breast cancer cell lines. Mir-31-5p has also been observed upregulated in Zinc Deficient rats compared to normal in ESCC and in other types of cancers when using this animal model. There has also been observed a strong encapsulation of tumour cells expressing miR-31, as well as a reduced cell survival rate. miR-31's antimetastatic effects therefore make it a potential therapeutic target for breast cancer. However, these two papers were formally retracted by the authors in 2015.

In molecular biology, mir-221 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.

miR-33 is a family of microRNA precursors, which are processed by the Dicer enzyme to give mature microRNAs. miR-33 is found in several animal species, including humans. In some species there is a single member of this family which gives the mature product mir-33. In humans there are two members of this family called mir-33a and mir-33b, which are located in intronic regions within two protein-coding genes for Sterol regulatory element-binding proteins respectively.
In molecular biology mir-365 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-370 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. This microRNA, mir-370-3p, has been shown to play a role in heart failure. The upregulation of mir-370-3p in the sinus node leads to downregulation of the pacemaker ion channel, HCN4, and thus downregulation of the corresponding ionic current, which causes sinus bradycardia.
In molecular biology mir-153 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-590 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-872 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-938 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.

MicroRNA 34a (miR-34a) is a MicroRNA that in humans is encoded by the MIR34A gene.













