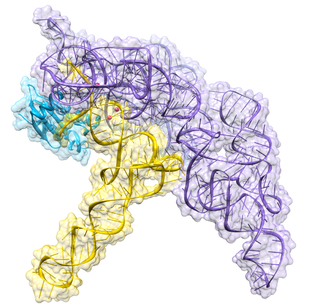
Ribonuclease P is a type of ribonuclease which cleaves RNA. RNase P is unique from other RNases in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way that a protein-based enzyme would. Its function is to cleave off an extra, or precursor, sequence of RNA on tRNA molecules. Further, RNase P is one of two known multiple turnover ribozymes in nature, the discovery of which earned Sidney Altman and Thomas Cech the Nobel Prize in Chemistry in 1989: in the 1970s, Altman discovered the existence of precursor tRNA with flanking sequences and was the first to characterize RNase P and its activity in processing of the 5' leader sequence of precursor tRNA. Recent findings also reveal that RNase P has a new function. It has been shown that human nuclear RNase P is required for the normal and efficient transcription of various small noncoding RNAs, such as tRNA, 5S rRNA, SRP RNA and U6 snRNA genes, which are transcribed by RNA polymerase III, one of three major nuclear RNA polymerases in human cells.

Polynucleotide Phosphorylase (PNPase) is a bifunctional enzyme with a phosphorolytic 3' to 5' exoribonuclease activity and a 3'-terminal oligonucleotide polymerase activity. That is, it dismantles the RNA chain starting at the 3' end and working toward the 5' end. It also synthesizes long, highly heteropolymeric tails in vivo. It accounts for all of the observed residual polyadenylation in strains of Escherichia coli missing the normal polyadenylation enzyme. Discovered by Marianne Grunberg-Manago working in Severo Ochoa's lab in 1955, the RNA-polymerization activity of PNPase was initially believed to be responsible for DNA-dependent synthesis of messenger RNA, a notion that was disproven by the late 1950s.

An exoribonuclease is an exonuclease ribonuclease, which are enzymes that degrade RNA by removing terminal nucleotides from either the 5' end or the 3' end of the RNA molecule. Enzymes that remove nucleotides from the 5' end are called 5'-3' exoribonucleases, and enzymes that remove nucleotides from the 3' end are called 3'-5' exoribonucleases.

Non-stop decay (NSD) is a cellular mechanism of mRNA surveillance to detect mRNA molecules lacking a stop codon and prevent these mRNAs from translation. The non-stop decay pathway releases ribosomes that have reached the far 3' end of an mRNA and guides the mRNA to the exosome complex, or to RNase R in bacteria for selective degradation. In contrast to nonsense-mediated decay (NMD), polypeptides do not release from the ribosome, and thus, NSD seems to involve mRNA decay factors distinct from NMD.
The Ski complex is a multi-protein complex involved in the 3' end degradation of messenger RNAs in yeast.

TRAMP complex is a multiprotein, heterotrimeric complex having distributive polyadenylation activity and identifies wide varieties of RNAs produced by polymerases. It was originally discovered in Saccharomycescerevisiae by LaCava et al., Vanacova et al. and Wyers et al. in 2005.

Exosome component 2, also known as EXOSC2, is a protein which in humans is encoded by the EXOSC2 gene.

Exosome component 10, also known as EXOSC10, is a human gene, the protein product of which is part of the exosome complex and is an autoantigen is patients with certain auto immune diseases, most notably scleromyositis.

Exosome complex exonuclease RRP44 or Dis3 is an enzyme that in humans is encoded by the DIS3 gene. Its protein product is an RNase enzyme homologous to the yeast protein Rrp44, and can be part of the exosome complex in the nucleus of eukaryotic cells.
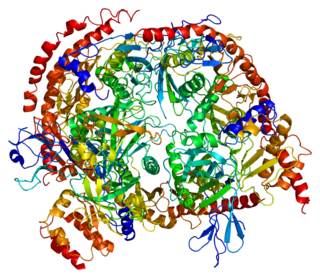
Exosome component 8, also known as EXOSC8, is a human gene, the protein product of which is part of the exosome complex.

Exosome component 7, also known as EXOSC7, is a human gene, the protein product of which is part of the exosome complex.

Exosome component 3, also known as EXOSC3, is a human gene, which is part of the exosome complex.

Exosome component 9, also known as EXOSC9, is a human gene, the protein product of which is part of the exosome complex and is an autoantigen is patients with certain auto immune diseases, most notably scleromyositis.

Exosome component 4, also known as EXOSC4, is a human gene, which is part of the exosome complex.

Superkiller viralicidic activity 2-like 2 is a protein that in humans is encoded by the SKIV2L2 gene.
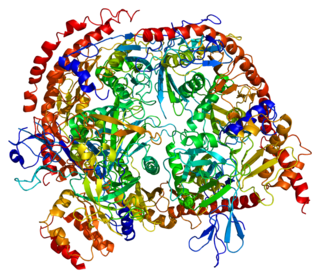
3'-5' exoribonuclease CSL4 homolog is an enzyme that in humans is encoded by the EXOSC1 gene.

Exosome component 5, also known as EXOSC5, is a human gene, which is part of the exosome complex.

Nuclear nucleic acid-binding protein C1D is a protein that in humans is encoded by the C1D gene. The C1D protein is encoded by a DNA binding gene traced in the nucleus. Protein C1D has a chromosomal location of 2p14. C1D has a family of proteins consisting of C1D homologues which may include Sas10 domains. C1D is thought to bind to RNA and DNA where it may be involved in mechanisms of DNA repair. Protein C1D is ubiquitously expressed in different human tissues.

Exosome complex exonuclease MTR3 is an enzyme that in humans is encoded by the EXOSC6 gene.
DIS3-like exonuclease 1 is an enzyme that in humans is encoded by the DIS3L gene. Its protein product is an RNase enzyme homologous to the yeast protein Rrp44, and can be part of the exosome complex in the cytoplasm of eukaryotic cells.

























