| mannitol-1-phosphate 5-dehydrogenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 1.1.1.17 | ||||||||
| CAS no. | 9028-24-4 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
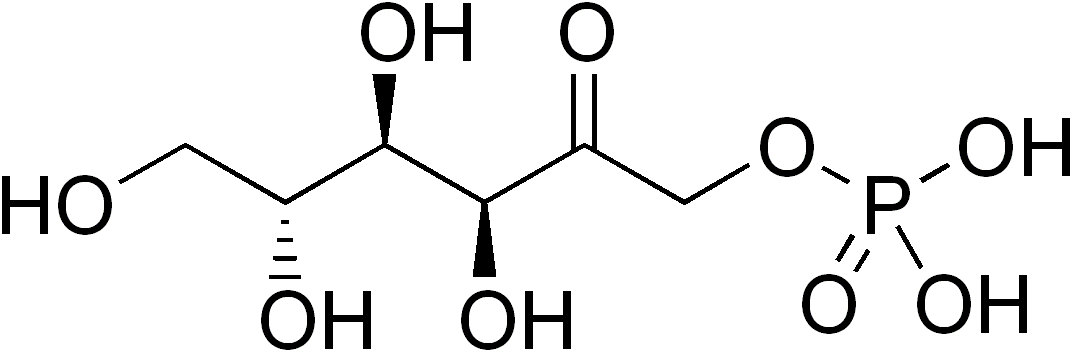
In enzymology, a mannitol-1-phosphate 5-dehydrogenase (EC 1.1.1.17) is an enzyme that catalyzes the chemical reaction
The two substrates of this enzyme are D-mannitol 1-phosphate and oxidised nicotinamide adenine dinucleotide (NAD+). Its products are fructose 6-phosphate, reduced NADH and a proton. [1] [2] [3] [4]
This enzyme belongs to the family of oxidoreductases, specifically those acting on the CH-OH group of donor with NAD+ or NADP+ as acceptor. The systematic name of this enzyme class is D-mannitol-1-phosphate:NAD+ 2-oxidoreductase. Other names in common use include hexose reductase, mannitol 1-phosphate dehydrogenase, D-mannitol-1-phosphate dehydrogenase, and fructose 6-phosphate reductase. This enzyme participates in fructose and mannose metabolism.