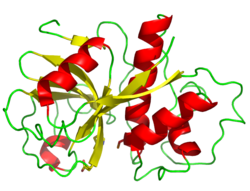
 Papain, the family's namesake member, from Carica papaya | |
| Identifiers | |
|---|---|
| Symbol | Peptidase_CA |
| Pfam clan | CL0125 |
| ECOD | 219.1.1 |
| MEROPS | CA |
Papain-like proteases (or papain-like (cysteine) peptidases; abbreviated PLP or PLCP) are a large protein family of cysteine protease enzymes that share structural and enzymatic properties with the group's namesake member, papain. They are found in all domains of life. In animals, the group is often known as cysteine cathepsins or, in older literature, lysosomal peptidases. [1] In the MEROPS protease enzyme classification system, papain-like proteases form Clan CA. [2] Papain-like proteases share a common catalytic dyad active site featuring a cysteine amino acid residue that acts as a nucleophile. [1]
Contents
The human genome encodes eleven cysteine cathepsins which have a broad range of physiological functions. [3] In some parasites papain-like proteases have roles in host invasion, such as cruzipain from Trypanosoma cruzi . [1] In plants, they are involved in host defense and in development. [4] Studies of papain-like proteases from prokaryotes have lagged their eukaryotic counterparts. [1] In cellular organisms they are synthesized as preproenzymes that are not enzymatically active until mature, and their activities are tightly regulated, often by the presence of endogenous protease inhibitors such as cystatins. [3] In many RNA viruses, including significant human pathogens such as the coronaviruses SARS-CoV and SARS-CoV-2, papain-like protease protein domains often have roles in processing of polyproteins into mature viral nonstructural proteins. [5] [6] Many papain-like proteases are considered potential drug targets. [3] [7]



