
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" base pairs allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins can recognize specific base-pairing patterns that identify particular regulatory regions of genes.

Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth.
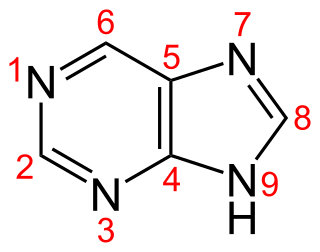
Purine is a heterocyclic aromatic organic compound that consists of two rings. It is water-soluble. Purine also gives its name to the wider class of molecules, purines, which include substituted purines and their tautomers. They are the most widely occurring nitrogen-containing heterocycles in nature.

Nucleobases, also known as nitrogenous bases or often simply bases, are nitrogen-containing biological compounds that form nucleosides, which, in turn, are components of nucleotides, with all of these monomers constituting the basic building blocks of nucleic acids. The ability of nucleobases to form base pairs and to stack one upon another leads directly to long-chain helical structures such as ribonucleic acid (RNA) and deoxyribonucleic acid (DNA).
A nucleoside triphosphate is a molecule containing a nitrogenous base bound to a 5-carbon sugar, with three phosphate groups bound to the sugar. It is an example of a nucleotide. They are the molecular precursors of both DNA and RNA, which are chains of nucleotides made through the processes of DNA replication and transcription. Nucleoside triphosphates also serve as a source of energy for cellular reactions and are involved in signalling pathways.
Hypoxanthine-guanine phosphoribosyltransferase (HGPRT) is an enzyme encoded in humans by the HPRT1 gene.

Inosinic acid or inosine monophosphate (IMP) is a nucleotide. Widely used as a flavor enhancer, it is typically obtained from chicken byproducts or other meat industry waste. Inosinic acid is important in metabolism. It is the ribonucleotide of hypoxanthine and the first nucleotide formed during the synthesis of purine nucleotides. It can also be formed by the deamination of adenosine monophosphate by AMP deaminase. It can be hydrolysed to inosine.
Purine nucleoside phosphorylase (PNP) also known as PNPase and inosine phosphorylase is an enzyme that in humans is encoded by the NP gene.

Nucleic acid metabolism is the process by which nucleic acids are synthesized and degraded. Nucleic acids are polymers of nucleotides. Nucleotide synthesis is an anabolic mechanism generally involving the chemical reaction of phosphate, pentose sugar, and a nitrogenous base. Destruction of nucleic acid is a catabolic reaction. Additionally, parts of the nucleotides or nucleobases can be salvaged to recreate new nucleotides. Both synthesis and degradation reactions require enzymes to facilitate the event. Defects or deficiencies in these enzymes can lead to a variety of diseases.
Purine metabolism refers to the metabolic pathways to synthesize and break down purines that are present in many organisms.

Xanthosine monophosphate is an intermediate in purine metabolism. It is a ribonucleoside monophosphate. It is formed from IMP via the action of IMP dehydrogenase, and it forms GMP via the action of GMP synthase. Also, XMP can be released from XTP by enzyme deoxyribonucleoside triphosphate pyrophosphohydrolase containing (d)XTPase activity.

Guanosine monophosphate synthetase, also known as GMPS is an enzyme that converts xanthosine monophosphate to guanosine monophosphate.

Amidophosphoribosyltransferase (ATase), also known as glutamine phosphoribosylpyrophosphate amidotransferase (GPAT), is an enzyme responsible for catalyzing the conversion of 5-phosphoribosyl-1-pyrophosphate (PRPP) into 5-phosphoribosyl-1-amine (PRA), using the amine group from a glutamine side-chain. This is the committing step in de novo purine synthesis. In humans it is encoded by the PPAT gene. ATase is a member of the purine/pyrimidine phosphoribosyltransferase family.
In enzymology, a nucleoside-triphosphate diphosphatase (EC 3.6.1.19) is an enzyme that catalyzes the chemical reaction

Inosine triphosphate pyrophosphatase is an enzyme that in humans is encoded by the ITPA gene, by the rdgB gene in bacteria E.coli and the HAM1 gene in yeast S. cerevisiae. Two transcript variants encoding two different isoforms have been found for this gene. Also, at least two other transcript variants have been identified which are probably regulatory rather than protein-coding.

Inosine-5′-monophosphate dehydrogenase (IMPDH) is a purine biosynthetic enzyme that catalyzes the nicotinamide adenine dinucleotide (NAD+)-dependent oxidation of inosine monophosphate (IMP) to xanthosine monophosphate (XMP), the first committed and rate-limiting step towards the de novo biosynthesis of guanine nucleotides from IMP. IMPDH is a regulator of the intracelluar guanine nucleotide pool, and is therefore important for DNA and RNA synthesis, signal transduction, energy transfer, glycoprotein synthesis, as well as other process that are involved in cellular proliferation.
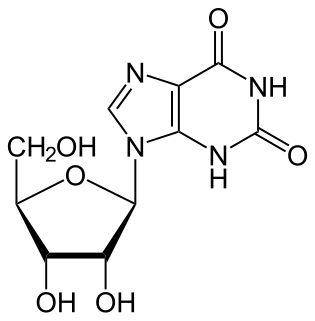
Xanthosine is a nucleoside derived from xanthine and ribose. It is the biosynthetic precursor to 7-methylxanthosine by the action of 7-methylxanthosine synthase. 7-Methylxanthosine in turn is the precursor to theobromine, which in turn is the precursor to caffeine, the alkaloid in coffee and tea.
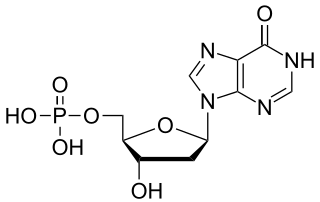
Deoxyinosine monophosphate (dIMP) is a nucleoside monophosphate and a derivative of inosinic acid. It can be formed by the deamination of the purine base in deoxyadenosine monophosphate (dAMP). The enzyme deoxyribonucleoside triphosphate pyrophosphohydrolase, encoded by YJR069C in S. cerevisiae and containing (d)ITPase and (d)XTPase activities, hydrolyses dITP, resulting in the release of pyrophosphate and dIMP.

Inosine triphosphate (ITP) is an intermediate in the purine metabolism pathway, seen in the synthesis of ATP and GTP. It comprises an inosine nucleotide containing three phosphate groups esterified to the sugar moiety.















