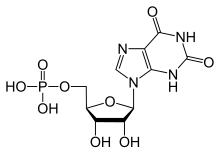
Nucleotides are organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver.
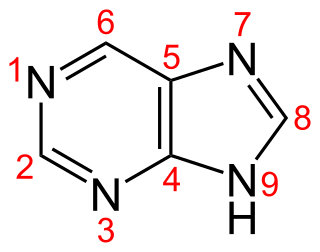
Purine is a heterocyclic aromatic organic compound that consists of two rings fused together. It is water-soluble. Purine also gives its name to the wider class of molecules, purines, which include substituted purines and their tautomers. They are the most widely occurring nitrogen-containing heterocycles in nature.

Uric acid is a heterocyclic compound of carbon, nitrogen, oxygen, and hydrogen with the formula C5H4N4O3. It forms ions and salts known as urates and acid urates, such as ammonium acid urate. Uric acid is a product of the metabolic breakdown of purine nucleotides, and it is a normal component of urine. High blood concentrations of uric acid can lead to gout and are associated with other medical conditions, including diabetes and the formation of ammonium acid urate kidney stones.
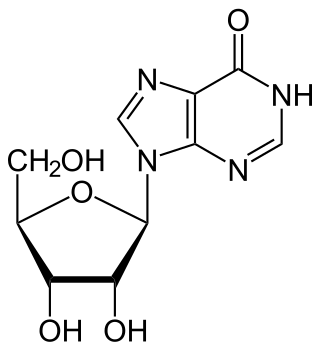
Inosine is a nucleoside that is formed when hypoxanthine is attached to a ribose ring (also known as a ribofuranose) via a β-N9-glycosidic bond. It was discovered in 1965 in analysis of RNA transferase. Inosine is commonly found in tRNAs and is essential for proper translation of the genetic code in wobble base pairs.

In biochemistry, a ribonucleotide is a nucleotide containing ribose as its pentose component. It is considered a molecular precursor of nucleic acids. Nucleotides are the basic building blocks of DNA and RNA. Ribonucleotides themselves are basic monomeric building blocks for RNA. Deoxyribonucleotides, formed by reducing ribonucleotides with the enzyme ribonucleotide reductase (RNR), are essential building blocks for DNA. There are several differences between DNA deoxyribonucleotides and RNA ribonucleotides. Successive nucleotides are linked together via phosphodiester bonds.
A salvage pathway is a pathway in which a biological product is produced from intermediates in the degradative pathway of its own or a similar substance. The term often refers to nucleotide salvage in particular, in which nucleotides are synthesized from intermediates in their degradative pathway.
A nucleoside triphosphate is a nucleoside containing a nitrogenous base bound to a 5-carbon sugar, with three phosphate groups bound to the sugar. They are the molecular precursors of both DNA and RNA, which are chains of nucleotides made through the processes of DNA replication and transcription. Nucleoside triphosphates also serve as a source of energy for cellular reactions and are involved in signalling pathways.

Hypoxanthine-guanine phosphoribosyltransferase (HGPRT) is an enzyme encoded in humans by the HPRT1 gene.

Inosinic acid or inosine monophosphate (IMP) is a nucleotide. Widely used as a flavor enhancer, it is typically obtained from chicken byproducts or other meat industry waste. Inosinic acid is important in metabolism. It is the ribonucleotide of hypoxanthine and the first nucleotide formed during the synthesis of purine nucleotides. It can also be formed by the deamination of adenosine monophosphate by AMP deaminase. It can be hydrolysed to inosine.
A nucleotidase is a hydrolytic enzyme that catalyzes the hydrolysis of a nucleotide into a nucleoside and a phosphate.

Nucleic acid metabolism is a collective term that refers to the variety of chemical reactions by which nucleic acids are either synthesized or degraded. Nucleic acids are polymers made up of a variety of monomers called nucleotides. Nucleotide synthesis is an anabolic mechanism generally involving the chemical reaction of phosphate, pentose sugar, and a nitrogenous base. Degradation of nucleic acids is a catabolic reaction and the resulting parts of the nucleotides or nucleobases can be salvaged to recreate new nucleotides. Both synthesis and degradation reactions require multiple enzymes to facilitate the event. Defects or deficiencies in these enzymes can lead to a variety of diseases.
Purine metabolism refers to the metabolic pathways to synthesize and break down purines that are present in many organisms.

Guanosine monophosphate synthetase, also known as GMPS is an enzyme that converts xanthosine monophosphate to guanosine monophosphate.
GMP reductase EC 1.7.1.7 is an enzyme that catalyzes the irreversible and NADPH-dependent reductive deamination of GMP into IMP.

Xanthosine 5'-triphosphate (XTP) is a nucleotide that is not produced by - and has no known function in - living cells. Uses of XTP are, in general, limited to experimental procedures on enzymes that bind other nucleotides. Deamination of purine bases can result in accumulation of such nucleotides as ITP, dITP, XTP, and dXTP.
In enzymology, a nucleoside-triphosphate diphosphatase (EC 3.6.1.19) is an enzyme that catalyzes the chemical reaction

Inosine-5′-monophosphate dehydrogenase (IMPDH) is a purine biosynthetic enzyme that catalyzes the nicotinamide adenine dinucleotide (NAD+)-dependent oxidation of inosine monophosphate (IMP) to xanthosine monophosphate (XMP), the first committed and rate-limiting step towards the de novo biosynthesis of guanine nucleotides from IMP. IMPDH is a regulator of the intracellular guanine nucleotide pool, and is therefore important for DNA and RNA synthesis, signal transduction, energy transfer, glycoprotein synthesis, as well as other process that are involved in cellular proliferation.
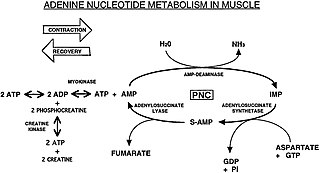
The Purine Nucleotide Cycle is a metabolic pathway in protein metabolism requiring the amino acids aspartate and glutamate. The cycle is used to regulate the levels of adenine nucleotides, in which ammonia and fumarate are generated. AMP coverts into IMP and the byproduct ammonia. IMP converts to S-AMP (adenylosuccinate), which then coverts to AMP and the byproduct fumarate. The fumarate goes on to produce ATP (energy) via oxidative phosphorylation as it enters the Krebs Cycle and then the Electron Transport Chain. Lowenstein first described this pathway and outlined its importance in processes including amino acid catabolism and regulation of flux through glycolysis and the Krebs cycle.
The gua operon is responsible for regulating the synthesis of guanosine mono phosphate (GMP), a purine nucleotide, from inosine monophosphate. It consists of two structural genes guaB (encodes for IMP dehydrogenase or and guaA apart from the promoter and operator region.
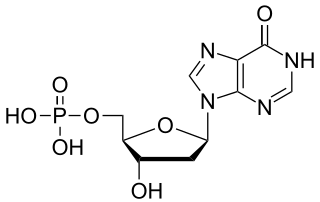
Deoxyinosine monophosphate (dIMP) is a nucleoside monophosphate and a derivative of inosinic acid. It can be formed by the deamination of the purine base in deoxyadenosine monophosphate (dAMP). The enzyme deoxyribonucleoside triphosphate pyrophosphohydrolase, encoded by YJR069C in S. cerevisiae and containing (d)ITPase and (d)XTPase activities, hydrolyses dITP, resulting in the release of pyrophosphate and dIMP.