Related Research Articles
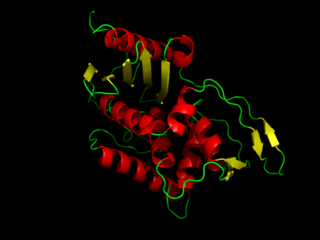
DD-transpeptidase is a bacterial enzyme that catalyzes the transfer of the R-L-αα-D-alanyl moiety of R-L-αα-D-alanyl-D-alanine carbonyl donors to the γ-OH of their active-site serine and from this to a final acceptor. It is involved in bacterial cell wall biosynthesis, namely, the transpeptidation that crosslinks the peptide side chains of peptidoglycan strands.

In enzymology, a D-alanine—D-alanine ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramate—L-alanine ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-L-alanine—D-glutamate ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-L-alanyl-D-glutamate—L-lysine ligase is an enzyme that catalyzes the chemical reaction
In enzymology, a UDP-N-acetylmuramoyl-tripeptide—D-alanyl-D-alanine ligase is an enzyme that catalyzes the chemical reaction
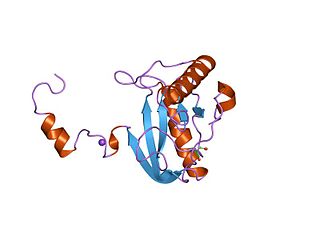
In enzymology, a N-acetylmuramoyl-L-alanine amidase is an enzyme that catalyzes a chemical reaction that cleaves the link between N-acetylmuramoyl residues and L-amino acid residues in certain cell-wall glycopeptides.
In enzymology, an UDP-N-acetylmuramoylpentapeptide-lysine N6-alanyltransferase (EC 2.3.2.10) is an enzyme that catalyzes the chemical reaction

Peptidoglycan binding domains have a general peptidoglycan binding function and a common core structure consisting of a closed, three-helical bundle with a left-handed twist. It is found at the N or C terminus of a variety of enzymes involved in bacterial cell wall degradation. Examples are:
The bacterial cell wall provides strength and rigidity to counteract internal osmotic pressure, and protection against the environment. The peptidoglycan layer gives the cell wall its strength, and helps maintain the overall shape of the cell. The basic peptidoglycan structure of both Gram-positive and Gram-negative bacteria comprises a sheet of glycan chains connected by short cross-linking polypeptides. Biosynthesis of peptidoglycan is a multi-step process comprising three main stages:
- formation of UDP-N-acetylmuramic acid (UDPMurNAc) from N-acetylglucosamine (GlcNAc).
- addition of a short polypeptide chain to the UDPMurNAc.
- addition of a second GlcNAc to the disaccharide-pentapeptide building block and transport of this unit through the cytoplasmic membrane and incorporation into the growing peptidoglycan layer.
In molecular biology, VanY are protein domains found in enzymes named metallopeptidases. They are vital to bacterial cell wall synthesis and antibiotic resistance.
Lipid II:glycine glycyltransferase (EC 2.3.2.16, N-acetylmuramoyl-L-alanyl-D-glutamyl-L-lysyl-D-alanyl-D-alanine-diphosphoundecaprenyl-N-acetylglucosamine:N6-glycine transferase, femX (gene)) is an enzyme with systematic name alanyl-D-alanine-diphospho-ditrans, octacis-undecaprenyl-N-acetylglucosamine:glycine N6-glycyltransferase. This enzyme catalyses the following chemical reaction
N-acetylmuramoyl-L-alanyl-D-glutamyl-L-lysyl-(N6-glycyl)-D-alanyl-D-alanine-diphosphoundecaprenyl-N-acetylglucosamine:glycine glycyltransferase (EC 2.3.2.17, femA (gene)) is an enzyme with systematic name N-acetylmuramoyl-L-alanyl-D-glutamyl-L-lysyl-(N6-glycyl)-D-alanyl-D-alanine-ditrans,octacis-diphosphoundecaprenyl-N-acetylglucosamine:glycine glycyltransferase. This enzyme catalyses the following chemical reaction
N-acetylmuramoyl-L-alanyl-D-glutamyl-L-lysyl-(N6-triglycine)-D-alanyl-D-alanine-diphosphoundecaprenyl-N-acetylglucosamine:glycine glycyltransferase (EC 2.3.2.18, femB (gene)) is an enzyme with systematic name N-acetylmuramoyl-L-alanyl-D-glutamyl-L-lysyl-(N6-triglycine)-D-alanyl-D-alanine-ditrans,octacis-diphosphoundecaprenyl-N-acetylglucosamine:glycine glycyltransferase. This enzyme catalyses the following chemical reaction
Alanine carboxypeptidase is an enzyme. This enzyme catalyses the following chemical reaction
Muramoyltetrapeptide carboxypeptidase is an enzyme. This enzyme catalyses the following chemical reaction
Zinc D-Ala-D-Ala carboxypeptidase (EC 3.4.17.14, Zn2+ G peptidase, D-alanyl-D-alanine hydrolase, D-alanyl-D-alanine-cleaving carboxypeptidase, DD-carboxypeptidase, G enzyme, DD-carboxypeptidase-transpeptidase) is an enzyme. This enzyme catalyses the following chemical reaction
UDP-N-acetylmuramoyl-L-alanyl-D-glutamate—2,6-diaminopimelate ligase is an enzyme with systematic name UDP-N-acetylmuramoyl-L-alanyl-D-glutamate:meso-2,6-diaminoheptanedioate gamma-ligase (ADP-forming). This enzyme catalyses the following chemical reaction
UDP-N-acetylmuramoyl-L-alanyl-D-glutamate—D-lysine ligase is an enzyme with systematic name UDP-N-acetylmuramoyl-L-alanyl-D-glutamate:D-lysine alpha-ligase (ADP-forming). This enzyme catalyses the following chemical reaction