
Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called proteases, but may also occur by intra-molecular digestion.
Protein targeting or protein sorting is the biological mechanism by which proteins are transported to their appropriate destinations within or outside the cell. Proteins can be targeted to the inner space of an organelle, different intracellular membranes, the plasma membrane, or to the exterior of the cell via secretion. Information contained in the protein itself directs this delivery process. Correct sorting is crucial for the cell; errors or dysfunction in sorting have been linked to multiple diseases.

A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism, and cell signaling.
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.
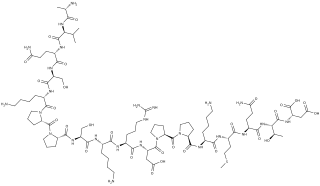
Systemin is a plant peptide hormone involved in the wound response in the family Solanaceae. It was the first plant hormone that was proven to be a peptide having been isolated from tomato leaves in 1991 by a group led by Clarence A. Ryan. Since then, other peptides with similar functions have been identified in tomato and outside of the Solanaceae. Hydroxyproline-rich glycopeptides were found in tobacco in 2001 and AtPeps were found in Arabidopsis thaliana in 2006. Their precursors are found both in the cytoplasm and cell walls of plant cells, upon insect damage, the precursors are processed to produce one or more mature peptides. The receptor for systemin was first thought to be the same as the brassinolide receptor but this is now uncertain. The signal transduction processes that occur after the peptides bind are similar to the cytokine-mediated inflammatory immune response in animals. Early experiments showed that systemin travelled around the plant after insects had damaged the plant, activating systemic acquired resistance, now it is thought that it increases the production of jasmonic acid causing the same result. The main function of systemins is to coordinate defensive responses against insect herbivores but they also affect plant development. Systemin induces the production of protease inhibitors which protect against insect herbivores, other peptides activate defensins and modify root growth. They have also been shown to affect plants' responses to salt stress and UV radiation. AtPEPs have been shown to affect resistance against oomycetes and may allow A. thaliana to distinguish between different pathogens. In Nicotiana attenuata, some of the peptides have stopped being involved in defensive roles and instead affect flower morphology.
In molecular biology, the Signal Peptide Peptidase (SPP) is a type of protein that specifically cleaves parts of other proteins. It is an intramembrane aspartyl protease with the conserved active site motifs 'YD' and 'GxGD' in adjacent transmembrane domains (TMDs). Its sequences is highly conserved in different vertebrate species. SPP cleaves remnant signal peptides left behind in membrane by the action of signal peptidase and also plays key roles in immune surveillance and the maturation of certain viral proteins.

Prolyl endopeptidase (PE) also known as prolyl oligopeptidase or post-proline cleaving enzyme is an enzyme that in humans is encoded by the PREP gene.

Thimet oligopeptidase is an enzyme that in humans is encoded by the THOP1 gene.

Neurolysin, mitochondrial is a protein that in humans is encoded by the NLN gene. It is a 78-kDa enzyme, widely distributed in mammalian tissues and found in various subcellular locations that vary with cell type. Neurolysin exemplifies the ability of neuropeptidases to target various cleavage site sequences by hydrolyzing them in vitro, and metabolism of neurotensin is the most important role of neurolysin in vivo. Neurolysin has also been implicated in pain control, blood pressure regulation, sepsis, reproduction, cancer biology pathogenesis of stroke, and glucose metabolism.

Puromycin-sensitive amino peptidase also known as cytosol alanyl aminopeptidase or alanine aminopeptidase (AAP) is an enzyme that in humans is encoded by the NPEPPS gene. It is used as a biomarker to detect damage to the kidneys, and that may be used to help diagnose certain kidney disorders. It is found at high levels in the urine when there are kidney problems.

Mitochondrial-processing peptidase subunit beta is an enzyme that in humans is encoded by the PMPCB gene. This gene is a member of the peptidase M16 family and encodes a protein with a zinc-binding motif. This protein is located in the mitochondrial matrix and catalyzes the cleavage of the leader peptides of precursor proteins newly imported into the mitochondria, though it only functions as part of a heterodimeric complex.

Mitochondrial-processing peptidase subunit alpha is an enzyme that in humans is encoded by the PMPCA gene. This gene PMPCA encoded a protein that is a member of the peptidase M16 family. This protein is located in the mitochondrial matrix and catalyzes the cleavage of the leader peptides of precursor proteins newly imported into the mitochondria, though it only functions as part of a heterodimeric complex.
Peptide signaling plays a significant role in various aspects of plant growth and development and specific receptors for various peptides have been identified as being membrane-localized receptor kinases, the largest family of receptor-like molecules in plants. Signaling peptides include members of the following protein families.
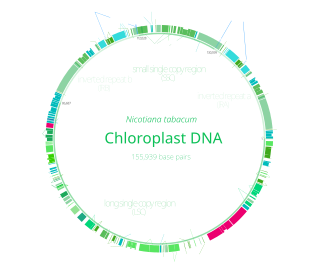
Chloroplast DNA (cpDNA) is the DNA located in chloroplasts, which are photosynthetic organelles located within the cells of some eukaryotic organisms. Chloroplasts, like other types of plastid, contain a genome separate from that in the cell nucleus. The existence of chloroplast DNA was identified biochemically in 1959, and confirmed by electron microscopy in 1962. The discoveries that the chloroplast contains ribosomes and performs protein synthesis revealed that the chloroplast is genetically semi-autonomous. The first complete chloroplast genome sequences were published in 1986, Nicotiana tabacum (tobacco) by Sugiura and colleagues and Marchantia polymorpha (liverwort) by Ozeki et al. Since then, a great number of chloroplast DNAs from various species have been sequenced.

An Oligopeptidase is an enzyme that cleaves peptides but not proteins. This property is due to its structure: the active site of this enzyme is located at the end of a narrow cavity which can only be reached by peptides.

Astacins are a family of multidomain metalloendopeptidases which are either secreted or membrane-anchored. These metallopeptidases belong to the MEROPS peptidase family M12, subfamily M12A. The protein fold of the peptidase domain for members of this family resembles that of thermolysin, the type example for clan MA and the predicted active site residues for members of this family and thermolysin occur in the motif HEXXH.
A target peptide is a short peptide chain that directs the transport of a protein to a specific region in the cell, including the nucleus, mitochondria, endoplasmic reticulum (ER), chloroplast, apoplast, peroxisome and plasma membrane. Some target peptides are cleaved from the protein by signal peptidases after the proteins are transported.
Caricain is an enzyme. This enzyme catalyses the following chemical reaction: Hydrolysis of proteins with broad specificity for peptide bonds, similar to those of papain and chymopapain
Chaperones, also called molecular chaperones, are proteins that assist other proteins in assuming their three-dimensional fold, which is necessary for protein function. However, the fold of a protein is sensitive to environmental conditions, such as temperature and pH, and thus chaperones are needed to keep proteins in their functional fold across various environmental conditions. Chaperones are an integral part of a cell's protein quality control network by assisting in protein folding and are ubiquitous across diverse biological taxa. Since protein folding, and therefore protein function, is susceptible to environmental conditions, chaperones could represent an important cellular aspect of biodiversity and environmental tolerance by organisms living in hazardous conditions. Chaperones also affect the evolution of proteins in general, as many proteins fundamentally require chaperones to fold or are naturally prone to misfolding, and therefore mitigates protein aggregation.













