Related Research Articles

HMG-CoA reductase is the rate-controlling enzyme of the mevalonate pathway, the metabolic pathway that produces cholesterol and other isoprenoids. HMGCR catalyzes the conversion of HMG-CoA to mevalonic acid, a necessary step in the biosynthesis of cholesterol. Normally in mammalian cells this enzyme is competitively suppressed so that its effect is controlled. This enzyme is the target of the widely available cholesterol-lowering drugs known collectively as the statins, which help treat dyslipidemia.

Michael Stuart Brown ForMemRS NAS AAA&S APS is an American geneticist and Nobel laureate. He was awarded the Nobel Prize in Physiology or Medicine with Joseph L. Goldstein in 1985 for describing the regulation of cholesterol metabolism.
In biochemistry, lipogenesis is the conversion of fatty acids and glycerol into fats, or a metabolic process through which acetyl-CoA is converted to triglyceride for storage in fat. Lipogenesis encompasses both fatty acid and triglyceride synthesis, with the latter being the process by which fatty acids are esterified to glycerol before being packaged into very-low-density lipoprotein (VLDL). Fatty acids are produced in the cytoplasm of cells by repeatedly adding two-carbon units to acetyl-CoA. Triacylglycerol synthesis, on the other hand, occurs in the endoplasmic reticulum membrane of cells by bonding three fatty acid molecules to a glycerol molecule. Both processes take place mainly in liver and adipose tissue. Nevertheless, it also occurs to some extent in other tissues such as the gut and kidney. A review on lipogenesis in the brain was published in 2008 by Lopez and Vidal-Puig. After being packaged into VLDL in the liver, the resulting lipoprotein is then secreted directly into the blood for delivery to peripheral tissues.

Sterol regulatory element-binding proteins (SREBPs) are transcription factors that bind to the sterol regulatory element DNA sequence TCACNCCAC. Mammalian SREBPs are encoded by the genes SREBF1 and SREBF2. SREBPs belong to the basic-helix-loop-helix leucine zipper class of transcription factors. Unactivated SREBPs are attached to the nuclear envelope and endoplasmic reticulum membranes. In cells with low levels of sterols, SREBPs are cleaved to a water-soluble N-terminal domain that is translocated to the nucleus. These activated SREBPs then bind to specific sterol regulatory element DNA sequences, thus upregulating the synthesis of enzymes involved in sterol biosynthesis. Sterols in turn inhibit the cleavage of SREBPs and therefore synthesis of additional sterols is reduced through a negative feed back loop.

Furin is a protease, a proteolytic enzyme activated by substrate presentation that in humans and other animals is encoded by the FURIN gene. Some proteins are inactive when they are first synthesized, and must have sections removed in order to become active. Furin cleaves these sections and activates the proteins. It was named furin because it was in the upstream region of an oncogene known as FES. The gene was known as FUR and therefore the protein was named furin. Furin is also known as PACE. A member of family S8, furin is a subtilisin-like peptidase.

ADAMs are a family of single-pass transmembrane and secreted metalloendopeptidases. All ADAMs are characterized by a particular domain organization featuring a pro-domain, a metalloprotease, a disintegrin, a cysteine-rich, an epidermal-growth factor like and a transmembrane domain, as well as a C-terminal cytoplasmic tail. Nonetheless, not all human ADAMs have a functional protease domain, which indicates that their biological function mainly depends on protein–protein interactions. Those ADAMs which are active proteases are classified as sheddases because they cut off or shed extracellular portions of transmembrane proteins. For example, ADAM10 can cut off part of the HER2 receptor, thereby activating it. ADAM genes are found in animals, choanoflagellates, fungi and some groups of green algae. Most green algae and all land plants likely lost ADAM proteins.
In molecular biology, the Signal Peptide Peptidase (SPP) is a type of protein that specifically cleaves parts of other proteins. It is an intramembrane aspartyl protease with the conserved active site motifs 'YD' and 'GxGD' in adjacent transmembrane domains (TMDs). Its sequences is highly conserved in different vertebrate species. SPP cleaves remnant signal peptides left behind in membrane by the action of signal peptidase and also plays key roles in immune surveillance and the maturation of certain viral proteins.
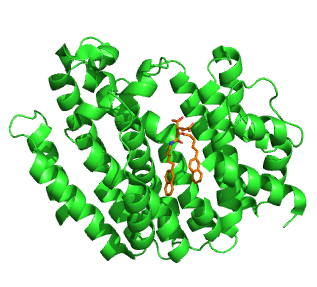
Squalene synthase (SQS) or farnesyl-diphosphate:farnesyl-diphosphate farnesyl transferase is an enzyme localized to the membrane of the endoplasmic reticulum. SQS participates in the isoprenoid biosynthetic pathway, catalyzing a two-step reaction in which two identical molecules of farnesyl pyrophosphate (FPP) are converted into squalene, with the consumption of NADPH. Catalysis by SQS is the first committed step in sterol synthesis, since the squalene produced is converted exclusively into various sterols, such as cholesterol, via a complex, multi-step pathway. SQS belongs to squalene/phytoene synthase family of proteins.

Sterol regulatory element-binding protein cleavage-activating protein, also known as SREBP cleavage-activating protein or SCAP, is a protein that in humans is encoded by the SCAP gene.
Xiaodong Wang is a Chinese-American biochemist best known for his work with apoptosis, one of the ways through which cells kill themselves.
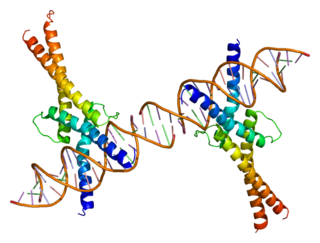
Sterol regulatory element-binding transcription factor 1 (SREBF1) also known as sterol regulatory element-binding protein 1 (SREBP-1) is a protein that in humans is encoded by the SREBF1 gene.

Sterol regulatory element-binding protein 2 (SREBP-2) also known as sterol regulatory element binding transcription factor 2 (SREBF2) is a protein that in humans is encoded by the SREBF2 gene.

Activating transcription factor 6, also known as ATF6, is a protein that, in humans, is encoded by the ATF6 gene and is involved in the unfolded protein response.

Membrane-bound transcription factor site-1 protease, or site-1 protease (S1P) for short, also known as subtilisin/kexin-isozyme 1 (SKI-1), is an enzyme that in humans is encoded by the MBTPS1 gene. S1P cleaves the endoplasmic reticulum loop of sterol regulatory element-binding protein (SREBP) transcription factors.

Insulin induced gene 1, also known as INSIG1, is a protein which in humans is encoded by the INSIG1 gene.

Oxysterol-binding protein 1 is a protein that in humans is encoded by the OSBP gene.

Insulin induced gene 2, also known as INSIG2, is a protein which in humans is encoded by the INSIG2 gene.

Presenilins-associated rhomboid-like protein, mitochondrial (PSARL), also known as PINK1/PGAM5-associated rhomboid-like protease (PARL), is an inner mitochondrial membrane protein that in humans is encoded by the PARL gene on chromosome 3. It is a member of the rhomboid family of intramembrane serine proteases. This protein is involved in signal transduction and apoptosis, as well as neurodegenerative diseases and type 2 diabetes.

The rhomboid proteases are a family of enzymes that exist in almost all species. They are proteases: they cut the polypeptide chain of other proteins. This proteolytic cleavage is irreversible in cells, and an important type of cellular regulation. Although proteases are one of the earliest and best studied class of enzyme, rhomboids belong to a much more recently discovered type: the intramembrane proteases. What is unique about intramembrane proteases is that their active sites are buried in the lipid bilayer of cell membranes, and they cleave other transmembrane proteins within their transmembrane domains. About 30% of all proteins have transmembrane domains, and their regulated processing often has major biological consequences. Accordingly, rhomboids regulate many important cellular processes, and may be involved in a wide range of human diseases.
Intramembrane proteases (IMPs), also known as intramembrane-cleaving proteases (I-CLiPs), are enzymes that have the property of cleaving transmembrane domains of integral membrane proteins. All known intramembrane proteases are themselves integral membrane proteins with multiple transmembrane domains, and they have their active sites buried within the lipid bilayer of cellular membranes. Intramembrane proteases are responsible for proteolytic cleavage in the cell signaling process known as regulated intramembrane proteolysis (RIP).
References
- ↑ Brown MS, Goldstein JL (September 1999). "A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood". Proceedings of the National Academy of Sciences of the United States of America. 96 (20): 11041–8. Bibcode:1999PNAS...9611041B. doi: 10.1073/pnas.96.20.11041 . PMC 34238 . PMID 10500120.
- ↑ Rawson RB, Zelenski NG, Nijhawan D, Ye J, Sakai J, Hasan MT, et al. (December 1997). "Complementation cloning of S2P, a gene encoding a putative metalloprotease required for intramembrane cleavage of SREBPs". Molecular Cell. 1 (1): 47–57. doi: 10.1016/S1097-2765(00)80006-4 . PMID 9659902.
- ↑ Brown MS, Ye J, Rawson RB, Goldstein JL (February 2000). "Regulated intramembrane proteolysis: a control mechanism conserved from bacteria to humans". Cell. 100 (4): 391–8. doi: 10.1016/S0092-8674(00)80675-3 . PMID 10693756.