
A microRNA is a small single-stranded non-coding RNA molecule found in plants, animals and some viruses, that functions in RNA silencing and post-transcriptional regulation of gene expression. miRNAs function via base-pairing with complementary sequences within mRNA molecules. As a result, these mRNA molecules are silenced, by one or more of the following processes: (1) Cleavage of the mRNA strand into two pieces, (2) Destabilization of the mRNA through shortening of its poly(A) tail, and (3) Less efficient translation of the mRNA into proteins by ribosomes.

In molecular biology lin-4 is a microRNA (miRNA) that was identified from a study of developmental timing in the nematode Caenorhabditis elegans. It was the first to be discovered of the miRNAs, a class of non-coding RNAs involved in gene regulation. miRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a 21 nucleotide product. The extents of the hairpin precursors are not generally known and are estimated based on hairpin prediction. The products are thought to have regulatory roles through complete or partial complementarity to mRNA. The lin-4 gene has been found to lie within a 4.11kb intron of a separate host gene.

The miR-103 microRNA precursor, is a short non-coding RNA gene involved in gene regulation. miR-103 and miR-107 have now been predicted or experimentally confirmed in human.
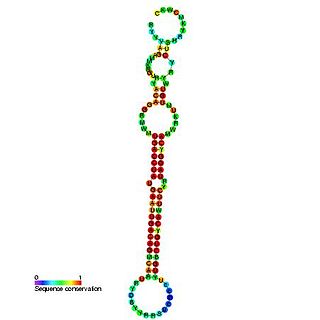
The miR-192 microRNA precursor, is a short non-coding RNA gene involved in gene regulation. miR-192 and miR-215 have now been predicted or experimentally confirmed in mouse and human.

In molecular biology, mir-46 and mir-47 are microRNA expressed in C. elegans from related hairpin precursor sequences. The predicted hairpin precursor sequences for Drosophila mir-281 are also related and, hence, belong to this family. The hairpin precursors are predicted based on base pairing and cross-species conservation; their extents are not known. In this case, the mature sequences are expressed from the 3' arms of the hairpin precursors.

miR-101 microRNA precursor is a small non-coding RNA that regulates gene expression. Expression of miR-101 has been validated in both human and mouse. This microRNA appears to be specific to the vertebrates and has now been predicted or confirmed in a wide range of vertebrate species. The precursor microRNA is a stem-loop structure of about 70 nucleotides in length that is processed by the Dicer enzyme to form the 21-24 nucleotide mature microRNA. In this case the mature sequence is excised from the 3' arm of the hairpin.

The miR-135 microRNA precursor is a small non-coding RNA that is involved in regulating gene expression. It has been shown to be expressed in human, mouse and rat. miR-135 has now been predicted or experimentally confirmed in a wide range of vertebrate species. Precursor microRNAs are ~70 nucleotides in length and are processed by the Dicer enzyme to produce the shorter 21-24 nucleotide mature sequence. In this case the mature sequence is excised from the 5' arm of the hairpin.

MicroRNA (miRNA) precursor miR156 is a family of plant non-coding RNA. This microRNA has now been predicted or experimentally confirmed in a range of plant species. Animal miRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. miR156 functions in the induction of flowering by suppressing the transcripts of SQUAMOSA-PROMOTER BINDING LIKE (SPL) transcription factors gene family. Its loading into ARGONAUTE1 and ARGONAUTE5 is required for full functionality in Arabidopsis thaliana. In plants the precursor sequences may be longer, and the carpel factory (caf) enzyme appears to be involved in processing. In this case the mature sequence comes from the 5' arm of the precursor, and both Arabidopsis thaliana and rice genomes contain a number of related miRNA precursors which give rise to almost identical mature sequences. The extents of the hairpin precursors are not generally known and are estimated based on hairpin prediction. The products are thought to have regulatory roles through complementarity to mRNA.
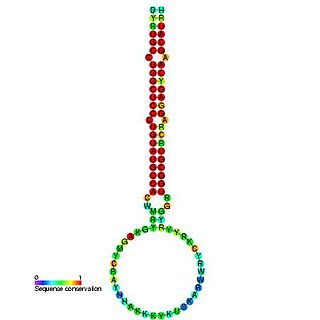
In molecular biology, mir-160 is a microRNA that has been predicted or experimentally confirmed in a range of plant species including Arabidopsis thaliana and Oryza sativa (rice). miR-160 is predicted to bind complementary sites in the untranslated regions of auxin response factor genes to regulate their expression. The hairpin precursors are predicted based on base pairing and cross-species conservation; their extents are not known. In this case, the mature sequence is excised from the 5' arm of the hairpin.

The mir-172 microRNA is thought to target mRNAs coding for APETALA2-like transcription factors. It has been verified experimentally in the model plant, Arabidopsis thaliana. The mature sequence is excised from the 3' arm of the hairpin.
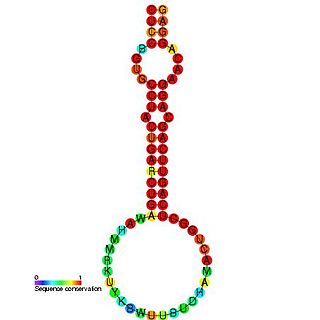
The miR-24 microRNA precursor is a small non-coding RNA molecule that regulates gene expression. microRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a mature ~22 nucleotide product. In this case the mature sequence comes from the 3' arm of the precursor. The mature products are thought to have regulatory roles through complementarity to mRNA. miR-24 is conserved in various species, and is clustered with miR-23 and miR-27, on human chromosome 9 and 19. Recently, miR-24 has been shown to suppress expression of two crucial cell cycle control genes, E2F2 and Myc in hematopoietic differentiation and also to promote keratinocyte differentiation by repressing actin-cytoskeleton regulators PAK4, Tsk5 and ArhGAP19.

The miR-29 microRNA precursor, or pre-miRNA, is a small RNA molecule in the shape of a stem-loop or hairpin. Each arm of the hairpin can be processed into one member of a closely related family of short non-coding RNAs that are involved in regulating gene expression. The processed, or "mature" products of the precursor molecule are known as microRNA (miRNA), and have been predicted or confirmed in a wide range of species.

The mir-2 microRNA family includes the microRNA genes mir-2 and mir-13. Mir-2 is widespread in invertebrates, and it is the largest family of microRNAs in the model species Drosophila melanogaster. MicroRNAs from this family are produced from the 3' arm of the precursor hairpin. Leaman et al. showed that the miR-2 family regulates cell survival by translational repression of proapoptotic factors. Based on computational prediction of targets, a role in neural development and maintenance has been suggested.

miR-30 microRNA precursor is a small non-coding RNA that regulates gene expression. Animal microRNAs are transcribed as pri-miRNA of varying length which in turns are processed in the nucleus by Drosha into ~70 nucleotide stem-loop precursor called pre-miRNA and subsequently processed by the Dicer enzyme to give a mature ~22 nucleotide product. In this case the mature sequence comes from both the 3' (miR-30) and 5' (mir-97-6) arms of the precursor. The products are thought to have regulatory roles through complementarity to mRNA.

mir-395 is a non-coding RNA called a microRNA that was identified in both Arabidopsis thaliana and Oryza sativa computationally and was later experimentally verified. mir-395 is thought to target mRNAs coding for ATP sulphurylases. The mature sequence is excised from the 3' arm of the hairpin.

mir-399 is a microRNA that was identified in both Arabidopsis thaliana and Oryza sativa computationally and was later experimentally verified. mir-399 is thought to target mRNAs coding for a phosphate transporter. The mature sequence is excised from the 3' arm of the hairpin. There are multiple copies of MIR399 in each plant genome, for example A. thaliana contains six microRNA precursors that all give rise to an almost identical mature miR-399 sequence.

The mir-6 microRNA precursor is a precursor microRNA specific to Drosophila species. In Drosophila melanogaster there are three mir-6 paralogs called dme-mir-6-1, dme-mir-6-2, dme-mir-6-3, which are clustered together in the genome. The extents of these hairpin precursors are estimated based on hairpin prediction. Each precursor is generated following the cleavage of a longer primary transcript in the nucleus, and is exported in the cytoplasm. In the cytoplasm, precursors are further processed by the enzyme Dicer, generating ~22 nucleotide products from each arm of the hairpin. The products generated from the 3' arm of each mir-6 precursor have identical sequences. Both 5' and 3' mature products are experimentally validated. Experimental data suggests that the mature products of mir-6 hairpins are expressed in the early embryo of Drosophila and target apoptotic genes such as hid, grim and rpr.
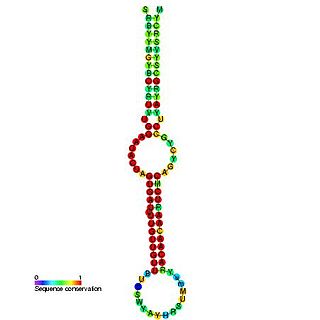
This family represents the microRNA (miRNA) precursor mir-7. This miRNA has been predicted or experimentally confirmed in a wide range of species. miRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. In this case the mature sequence comes from the 5' arm of the precursor. The extents of the hairpin precursors are not generally known and are estimated based on hairpin prediction. The involvement of Dicer in miRNA processing suggests a relationship with the phenomenon of RNA interference.

miR-96 microRNA precursor is a small non-coding RNA that regulates gene expression. microRNAs are transcribed as ~80 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~23 nucleotide products. In this case the mature sequence comes from the 5′ arm of the precursor. The mature products are thought to have regulatory roles through complementarity to mRNA.

miR-338 is a family of brain-specific microRNA precursors found in mammals, including humans. The ~22 nucleotide mature miRNA sequence is excised from the precursor hairpin by the enzyme Dicer. This sequence then associates with RISC which effects RNA interference.




















