
The miR-9 microRNA, is a short non-coding RNA gene involved in gene regulation. The mature ~21nt miRNAs are processed from hairpin precursor sequences by the Dicer enzyme. The dominant mature miRNA sequence is processed from the 5' arm of the mir-9 precursor, and from the 3' arm of the mir-79 precursor. The mature products are thought to have regulatory roles through complementarity to mRNA. In vertebrates, miR-9 is highly expressed in the brain, and is suggested to regulate neuronal differentiation. A number of specific targets of miR-9 have been proposed, including the transcription factor REST and its partner CoREST.
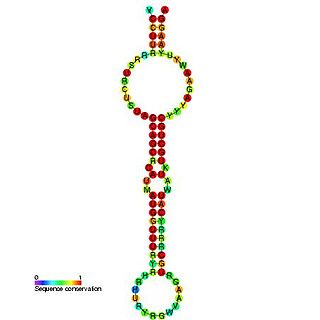
The miR-15 microRNA precursor family is made up of small non-coding RNA genes that regulate gene expression. The family includes the related mir-15a and mir-15b sequences, as well as miR-16-1, miR-16-2, miR-195 and miR-497. These six highly conserved miRNAs are clustered on three separate chromosomes. In humans miR-15a and miR-16 are clustered within 0.5 kilobases at chromosome position 13q14. This region has been found to be the most commonly affected in chronic lymphocytic leukaemia (CLL), with deletions of the entire region in more than half of cases. Both miR-15a and miR-16 are thus frequently deleted or down-regulated in CLL samples with 13q14 deletions; occurring in more than two thirds of CLL cases. The expression of miR-15a is associated with survival in triple negative breast cancer.
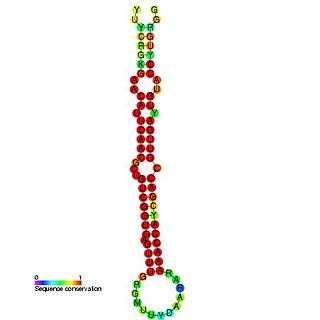
In molecular biology miR-181 microRNA precursor is a small non-coding RNA molecule. MicroRNAs (miRNAs) are transcribed as ~70 nucleotide precursors and subsequently processed by the RNase-III type enzyme Dicer to give a ~22 nucleotide mature product. In this case the mature sequence comes from the 5' arm of the precursor. They target and modulate protein expression by inhibiting translation and / or inducing degradation of target messenger RNAs. This new class of genes has recently been shown to play a central role in malignant transformation. miRNA are downregulated in many tumors and thus appear to function as tumor suppressor genes. The mature products miR-181a, miR-181b, miR-181c or miR-181d are thought to have regulatory roles at posttranscriptional level, through complementarity to target mRNAs. miR-181 has been predicted or experimentally confirmed in a wide number of vertebrate species such as rat, zebrafish, and pufferfish.

miR-196 is a non-coding RNA called a microRNA that has been shown to be expressed in humans and mice. miR-196 appears to be a vertebrate specific microRNA and has now been predicted or experimentally confirmed in a wide range of vertebrate species. In many species the miRNA appears to be expressed from intergenic regions in HOX gene clusters. The hairpin precursors are predicted based on base pairing and cross-species conservation—their extents are not known. In this case the mature sequence is excised from the 5' arm of the hairpin.

There are 89 known sequences today in the microRNA 19 (miR-19) family but it will change quickly. They are found in a large number of vertebrate species. The miR-19 microRNA precursor is a small non-coding RNA molecule that regulates gene expression. Within the human and mouse genome there are three copies of this microRNA that are processed from multiple predicted precursor hairpins:

The miR-1 microRNA precursor is a small micro RNA that regulates its target protein's expression in the cell. microRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give products at ~22 nucleotides. In this case the mature sequence comes from the 3' arm of the precursor. The mature products are thought to have regulatory roles through complementarity to mRNA. In humans there are two distinct microRNAs that share an identical mature sequence, and these are called miR-1-1 and miR-1-2.

The miR-29 microRNA precursor, or pre-miRNA, is a small RNA molecule in the shape of a stem-loop or hairpin. Each arm of the hairpin can be processed into one member of a closely related family of short non-coding RNAs that are involved in regulating gene expression. The processed, or "mature" products of the precursor molecule are known as microRNA (miRNA), and have been predicted or confirmed in a wide range of species.

The mir-2 microRNA family includes the microRNA genes mir-2 and mir-13. Mir-2 is widespread in invertebrates, and it is the largest family of microRNAs in the model species Drosophila melanogaster. MicroRNAs from this family are produced from the 3' arm of the precursor hairpin. Leaman et al. showed that the miR-2 family regulates cell survival by translational repression of proapoptotic factors. Based on computational prediction of targets, a role in neural development and maintenance has been suggested.
The miR-34 microRNA precursor family are non-coding RNA molecules that, in mammals, give rise to three major mature miRNAs. The miR-34 family members were discovered computationally and later verified experimentally. The precursor miRNA stem-loop is processed in the cytoplasm of the cell, with the predominant miR-34 mature sequence excised from the 5' arm of the hairpin.

The mir-6 microRNA precursor is a precursor microRNA specific to Drosophila species. In Drosophila melanogaster there are three mir-6 paralogs called dme-mir-6-1, dme-mir-6-2, dme-mir-6-3, which are clustered together in the genome. The extents of these hairpin precursors are estimated based on hairpin prediction. Each precursor is generated following the cleavage of a longer primary transcript in the nucleus, and is exported in the cytoplasm. In the cytoplasm, precursors are further processed by the enzyme Dicer, generating ~22 nucleotide products from each arm of the hairpin. The products generated from the 3' arm of each mir-6 precursor have identical sequences. Both 5' and 3' mature products are experimentally validated. Experimental data suggests that the mature products of mir-6 hairpins are expressed in the early embryo of Drosophila and target apoptotic genes such as hid, grim and rpr.
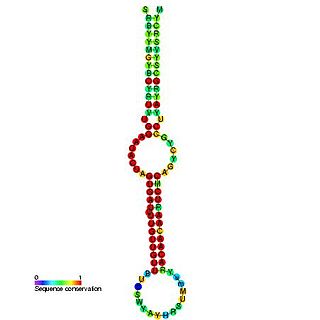
This family represents the microRNA (miRNA) precursor mir-7. This miRNA has been predicted or experimentally confirmed in a wide range of species. miRNAs are transcribed as ~70 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~22 nucleotide product. In this case the mature sequence comes from the 5' arm of the precursor. The extents of the hairpin precursors are not generally known and are estimated based on hairpin prediction. The involvement of Dicer in miRNA processing suggests a relationship with the phenomenon of RNA interference.

The miR-92 microRNAs are short single stranded non-protein coding RNA fragments initially discovered incorporated into an RNP complex with a proposed role of processing RNA molecules and further RNP assembly. Mir-92 has been mapped to the human genome as part of a larger cluster at chromosome 13q31.3, where it is 22 nucleotides in length but exists in the genome as part of a longer precursor sequence. There is an exact replica of the mir-92 precursor on the X chromosome. MicroRNAs are endogenous triggers of the RNAi pathway which involves several ribonucleic proteins (RNPs) dedicated to repressing mRNA molecules via translation inhibition and/or induction of mRNA cleavage. miRNAs are themselves matured from their long RNA precursors by ribonucleic proteins as part of a 2 step biogenesis mechanism involving RNA polymerase 2.

Homeobox D10, also known as HOXD10, is a protein which in humans is encoded by the HOXD10 gene.

In molecular biology mir-126 is a short non-coding RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several pre- and post-transcription mechanisms.

In molecular biology, mir-145 microRNA is a short RNA molecule that in humans is encoded by the MIR145 gene. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.

In molecular biology, miR-184 microRNA is a short non-coding RNA molecule. MicroRNAs (miRNAs) function as posttranscriptional regulators of expression levels of other genes by several mechanisms. Several targets for miR-184 have been described, including that of mediators of neurological development, apoptosis and it has been suggested that miR-184 plays an essential role in development.

In molecular biology MicroRNA-223 (miR-223) is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. miR-223 is a hematopoietic specific microRNA with crucial functions in myeloid lineage development. It plays an essential role in promoting granulocytic differentiation while also being associated with the suppression of erythrocytic differentiation. miR-223 is commonly repressed in hepatocellular carcinoma and leukemia. Higher expression levels of miRNA-223 are associated with extranodal marginal-zone lymphoma of mucosa-associated lymphoid tissue of the stomach and recurrent ovarian cancer. In some cancers the microRNA-223 down-regulation is correlated with higher tumor burden, disease aggressiveness, and poor prognostic factors. MicroRNA-223 is also associated with rheumatoid arthritis, sepsis, type 2 diabetes, and hepatic ischemia.
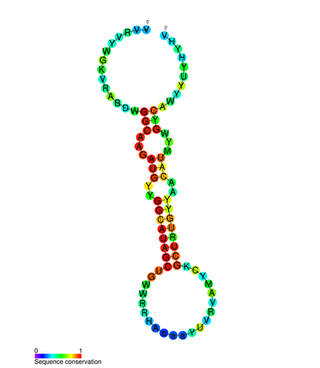
miR-31 has been characterised as a tumour suppressor miRNA, with its levels varying in breast cancer cells according to the metastatic state of the tumour. From its typical abundance in healthy tissue is a moderate decrease in non-metastatic breast cancer cell lines, and levels are almost completely absent in mouse and human metastatic breast cancer cell lines. Mir-31-5p has also been observed upregulated in Zinc Deficient rats compared to normal in ESCC and in other types of cancers when using this animal model. There has also been observed a strong encapsulation of tumour cells expressing miR-31, as well as a reduced cell survival rate. miR-31's antimetastatic effects therefore make it a potential therapeutic target for breast cancer. However, these two papers were formally retracted by the authors in 2015.

miR-191 is a family of microRNA precursors found in mammals, including humans. The ~22 nucleotide mature miRNA sequence is excised from the precursor hairpin by the enzyme Dicer. This sequence then associates with RISC which effects RNA interference.
mir-615 microRNA is a short non-coding RNA molecule belonging both to the family of microRNAs and to that of small interfering RNAs (siRNAs). MicroRNAs function to regulate the expression levels of other genes by several mechanisms, whilst siRNAs are involved primarily with the RNA interference (RNAi) pathway. siRNAs have been linked through some members to the regulation of cancer cell growth, specifically in prostate adenocarcinoma.


















