
Chronic lymphocytic leukemia (CLL) is a type of cancer in which the bone marrow makes too many lymphocytes. Early on, there are typically no symptoms. Later, non-painful lymph node swelling, feeling tired, fever, night sweats, or weight loss for no clear reason may occur. Enlargement of the spleen and low red blood cells (anemia) may also occur. It typically worsens gradually over years.

The mir-10 microRNA precursor is a short non-coding RNA gene involved in gene regulation. It is part of an RNA gene family which contains mir-10, mir-51, mir-57, mir-99 and mir-100. mir-10, mir-99 and mir-100 have now been predicted or experimentally confirmed in a wide range of species. miR-51 and miR-57 have currently only been identified in the nematode Caenorhabditis elegans.
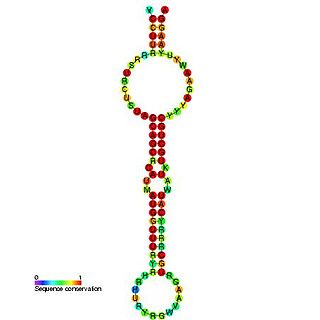
The miR-15 microRNA precursor family is made up of small non-coding RNA genes that regulate gene expression. The family includes the related mir-15a and mir-15b sequences, as well as miR-16-1, miR-16-2, miR-195 and miR-497. These six highly conserved miRNAs are clustered on three separate chromosomes. In humans miR-15a and miR-16 are clustered within 0.5 kilobases at chromosome position 13q14. This region has been found to be the most commonly affected in chronic lymphocytic leukaemia (CLL), with deletions of the entire region in more than half of cases. Both miR-15a and miR-16 are thus frequently deleted or down-regulated in CLL samples with 13q14 deletions; occurring in more than two thirds of CLL cases. The expression of miR-15a is associated with survival in triple negative breast cancer.
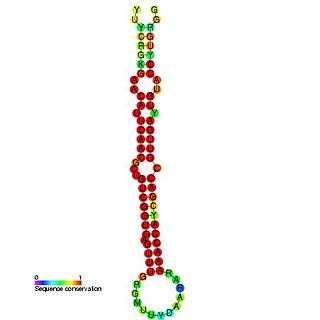
In molecular biology miR-181 microRNA precursor is a small non-coding RNA molecule. MicroRNAs (miRNAs) are transcribed as ~70 nucleotide precursors and subsequently processed by the RNase-III type enzyme Dicer to give a ~22 nucleotide mature product. In this case the mature sequence comes from the 5' arm of the precursor. They target and modulate protein expression by inhibiting translation and / or inducing degradation of target messenger RNAs. This new class of genes has recently been shown to play a central role in malignant transformation. miRNA are downregulated in many tumors and thus appear to function as tumor suppressor genes. The mature products miR-181a, miR-181b, miR-181c or miR-181d are thought to have regulatory roles at posttranscriptional level, through complementarity to target mRNAs. miR-181 which has been predicted or experimentally confirmed in a wide number of vertebrate species as rat, zebrafish, and in the pufferfish.

The miR-29 microRNA precursor, or pre-miRNA, is a small RNA molecule in the shape of a stem-loop or hairpin. Each arm of the hairpin can be processed into one member of a closely related family of short non-coding RNAs that are involved in regulating gene expression. The processed, or "mature" products of the precursor molecule are known as microRNA (miRNA), and have been predicted or confirmed in a wide range of species.
RCC1 and BTB domain-containing protein 1 is a protein that in humans is encoded by the RCBTB1 gene.

SPRY domain-containing protein 7 (SPRYD7) also known as chronic lymphocytic leukemia deletion region gene 6 protein (CLLD6) is a protein that in humans is encoded by the SPRYD7 gene.
Deleted in lymphocytic leukemia 2 is a long non-coding RNA that in humans is encoded by the DLEU2 gene. In humans it is located on chromosome 13q14. The DLEU2 gene was originally identified as a potential tumour suppressor gene and is often deleted in patients with B-cell chronic lymphocytic leukemia.
An oncomir is a microRNA (miRNA) that is associated with cancer. MicroRNAs are short RNA molecules about 22 nucleotides in length. Essentially, miRNAs specifically target certain messenger RNAs (mRNAs) to prevent them from coding for a specific protein. The dysregulation of certain microRNAs (oncomirs) has been associated with specific cancer forming (oncogenic) events. Many different oncomirs have been identified in numerous types of human cancers.

In molecular biology, mir-145 microRNA is a short RNA molecule that in humans is encoded by the MIR145 gene. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.

In molecular biology MicroRNA-223 (miR-223) is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms. miR-223 is a hematopoietic specific microRNA with crucial functions in myeloid lineage development. It plays an essential role in promoting granulocytic differentiation while also being associated with the suppression of erythrocytic differentiation. miR-223 is commonly repressed in hepatocellular carcinoma and leukemia. Higher expression levels of miRNA-223 are associated with extranodal marginal-zone lymphoma of mucosa-associated lymphoid tissue of the stomach and recurrent ovarian cancer. In some cancers the microRNA-223 down-regulation is correlated with higher tumor burden, disease aggressiveness, and poor prognostic factors. MicroRNA-223 is also associated with rheumatoid arthritis, sepsis, type 2 diabetes, and hepatic ischemia.

In molecular biology, mir-221 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
UCbase is a database of ultraconserved sequences that were first described by Bejerano, G. et al. in 2004. They are highly conserved genome regions that share 100% identity among human, mouse and rat. UCRs are 481 sequences longer than 200 bases. They are frequently located at genomic regions involved in cancer, differentially expressed in human leukemias and carcinomas and in some instances regulated by microRNAs. The first release of UCbase was published by Taccioli, C. et al. in 2009. Recent updates include new annotation based on hg19 Human genome, information about disorders related to the chromosome coordinates using the SNOMED CT classification, a query tool to search for SNPs, and a new text box to directly interrogate the database using a MySQL interface. Moreover, a sequence comparison tool allows the researchers to match selected sequences against ultraconserved elements located in genomic regions involved in specific disorders. To facilitate the interactive, visual interpretation of UCR chromosomal coordinates, the authors have implemented the graph visualization feature of UCbase creating a link to the UCSC Genome Browser. UCbase 2.0 does not provide microRNAs (miRNAs) information anymore focusing only on UCRs. The official release of UCbase 2.0 was published in 2014.

In molecular biology, deleted in lymphocytic leukemia 1 , also known as DLEU1, is a long non-coding RNA. In humans it is located on chromosome 13q14. The DLEU1 gene was originally identified as a potential tumour suppressor gene and is often deleted in patients with B-cell chronic lymphocytic leukemia. It was later discovered to be a long non-coding RNA with over 20 different splice variants.

miR-150 is a family of microRNA precursors found in mammals, including humans. The ~22 nucleotide mature miRNA sequence is excised from the precursor hairpin by the enzyme Dicer. This sequence then associates with RISC which effects RNA interference.

miR-191 is a family of microRNA precursors found in mammals, including humans. The ~22 nucleotide mature miRNA sequence is excised from the precursor hairpin by the enzyme Dicer. This sequence then associates with RISC which effects RNA interference.
MicroRNA sequencing (miRNA-seq), a type of RNA-Seq, is the use of next-generation sequencing or massively parallel high-throughput DNA sequencing to sequence microRNAs, also called miRNAs. miRNA-seq differs from other forms of RNA-seq in that input material is often enriched for small RNAs. miRNA-seq allows researchers to examine tissue-specific expression patterns, disease associations, and isoforms of miRNAs, and to discover previously uncharacterized miRNAs. Evidence that dysregulated miRNAs play a role in diseases such as cancer has positioned miRNA-seq to potentially become an important tool in the future for diagnostics and prognostics as costs continue to decrease. Like other miRNA profiling technologies, miRNA-Seq has both advantages and disadvantages.
In molecular biology mir-32 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-331 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.
In molecular biology mir-28 microRNA is a short RNA molecule. MicroRNAs function to regulate the expression levels of other genes by several mechanisms.












