| Avogadro | |
|---|---|
 Avogadro logo | |
| Initial release | February 29, 2008 |
| Stable release | 1.102.1 / 27 October 2025 |
| Repository | github |
| Written in | C++ (Qt) |
| Operating system | Linux, macOS, Unix, Windows |
| Platform | IA-32, x86-64 |
| Size | 85.3 MB |
| Available in | 14 languages |
List of languages Chinese, English, French, Hungarian, German, Indonesian, Japanese, Korean, Portuguese, Romanian, Spanish, Tamil, Turkish, Ukrainian | |
| Type | Molecule editor |
| License | GPL v2 |
| Website | avogadro |
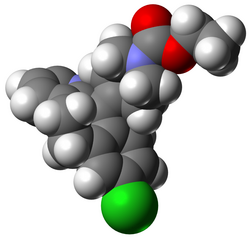
Avogadro is a molecule editor and visualizer designed for cross-platform use in computational chemistry, molecular modeling, bioinformatics, materials science, and related areas. [1] [2] [3] [4] It is extensible via a plugin architecture. [5]