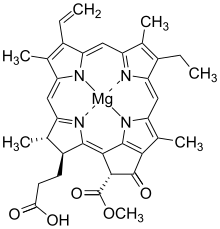
 Chlorophyllide a | |
| Names | |
|---|---|
| IUPAC name Magnesium (3S,4S,21R)-3-(2-carboxyethyl)-14-ethyl-21-(methoxycarbonyl)-4,8,13,18-tetramethyl-20-oxo-9-vinyl-23,25-didehydrophorbine-23,25-diide | |
| Identifiers | |
| |
| |
3D model (JSmol) | |
| ChEBI |
|
| ChemSpider | |
PubChem CID | |
| |
| |
| Properties | |
| C35H34MgN4O5 | |
| Molar mass | 614.973 g/mol |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa). | |
Chlorophyllide a and chlorophyllide b are the biosynthetic precursors of chlorophyll a and chlorophyll b respectively. Their propionic acid groups are converted to phytyl esters by the enzyme chlorophyll synthase in the final step of the pathway. Thus the main interest in these chemical compounds has been in the study of chlorophyll biosynthesis in plants, algae and cyanobacteria. Chlorophyllide a is also an intermediate in the biosynthesis of bacteriochlorophylls. [1] [2]
Contents
- Structures
- Biosynthesis steps up to formation of protoporphyrin IX
- Biosynthesis of chlorophyllides from protoporphyrin IX
- Insertion of magnesium
- Esterification of the ring C propionate group
- From porphyrin to chlorin
- Reduction steps to chlorophyllide a
- From chlorophyllide a to chlorophyllide b
- Use in the biosynthesis of chlorophylls
- Use in the biosynthesis of bacteriochlorophylls
- BChl a: bacteriochlorin ring and sidechains
- References






