Related Research Articles

Histone acetyltransferases (HATs) are enzymes that acetylate conserved lysine amino acids on histone proteins by transferring an acetyl group from acetyl-CoA to form ε-N-acetyllysine. DNA is wrapped around histones, and, by transferring an acetyl group to the histones, genes can be turned on and off. In general, histone acetylation increases gene expression.

Histone deacetylases (EC 3.5.1.98, HDAC) are a class of enzymes that remove acetyl groups (O=C-CH3) from an ε-N-acetyl lysine amino acid on both histone and non-histone proteins. HDACs allow histones to wrap the DNA more tightly. This is important because DNA is wrapped around histones, and DNA expression is regulated by acetylation and de-acetylation. HDAC's action is opposite to that of histone acetyltransferase. HDAC proteins are now also called lysine deacetylases (KDAC), to describe their function rather than their target, which also includes non-histone proteins. In general, they suppress gene expression.
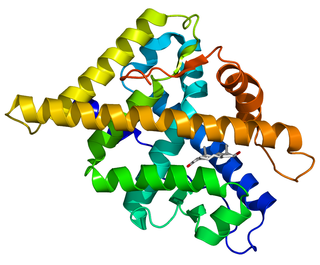
The androgen receptor (AR), also known as NR3C4, is a type of nuclear receptor that is activated by binding any of the androgenic hormones, including testosterone and dihydrotestosterone, in the cytoplasm and then translocating into the nucleus. The androgen receptor is most closely related to the progesterone receptor, and progestins in higher dosages can block the androgen receptor.
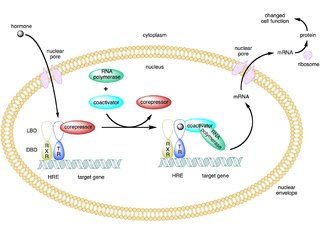
A coactivator is a type of transcriptional coregulator that binds to an activator to increase the rate of transcription of a gene or set of genes. The activator contains a DNA binding domain that binds either to a DNA promoter site or a specific DNA regulatory sequence called an enhancer. Binding of the activator-coactivator complex increases the speed of transcription by recruiting general transcription machinery to the promoter, therefore increasing gene expression. The use of activators and coactivators allows for highly specific expression of certain genes depending on cell type and developmental stage.
In molecular genetics, the Krüppel-like family of transcription factors (KLFs) are a set of eukaryotic C2H2 zinc finger DNA-binding proteins that regulate gene expression. This family has been expanded to also include the Sp transcription factor and related proteins, forming the Sp/KLF family.

Proline-, glutamic acid- and leucine-rich protein 1 (PELP1) also known as modulator of non-genomic activity of estrogen receptor (MNAR) and transcription factor HMX3 is a protein that in humans is encoded by the PELP1 gene. is a transcriptional corepressor for nuclear receptors such as glucocorticoid receptors and a coactivator for estrogen receptors.
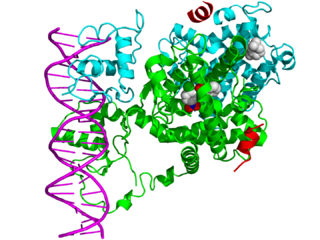
In the field of molecular biology, nuclear receptors are a class of proteins responsible for sensing steroids, thyroid hormones, vitamins, and certain other molecules. These intracellular receptors work with other proteins to regulate the expression of specific genes thereby controlling the development, homeostasis, and metabolism of the organism.

Histone acetylation and deacetylation are the processes by which the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are acetylated and deacetylated as part of gene regulation.

Histone deacetylase 1 (HDAC1) is an enzyme that in humans is encoded by the HDAC1 gene.

The nuclear receptor co-repressor 1 also known as thyroid-hormone- and retinoic-acid-receptor-associated co-repressor 1 (TRAC-1) is a protein that in humans is encoded by the NCOR1 gene.

The nuclear receptor co-repressor 2 (NCOR2) is a transcriptional coregulatory protein that contains several nuclear receptor-interacting domains. In addition, NCOR2 appears to recruit histone deacetylases to DNA promoter regions. Hence NCOR2 assists nuclear receptors in the down regulation of target gene expression. NCOR2 is also referred to as a silencing mediator for retinoid or thyroid-hormone receptors (SMRT) or T3 receptor-associating cofactor 1 (TRAC-1).

In genetics and molecular biology, a corepressor is a molecule that represses the expression of genes. In prokaryotes, corepressors are small molecules whereas in eukaryotes, corepressors are proteins. A corepressor does not directly bind to DNA, but instead indirectly regulates gene expression by binding to repressors.

Histone deacetylase 3 is an enzyme encoded by the HDAC3 gene in both humans and mice.

Paired amphipathic helix protein Sin3a is a protein that in humans is encoded by the SIN3A gene.

Tripartite motif-containing 28 (TRIM28), also known as transcriptional intermediary factor 1β (TIF1β) and KAP1, is a protein that in humans is encoded by the TRIM28 gene.

Metastasis-associated protein MTA1 is a protein that in humans is encoded by the MTA1 gene. MTA1 is the founding member of the MTA family of genes. MTA1 is primarily localized in the nucleus but also found to be distributed in the extra-nuclear compartments. MTA1 is a component of several chromatin remodeling complexes including the nucleosome remodeling and deacetylation complex (NuRD). MTA1 regulates gene expression by functioning as a coregulator to integrate DNA-interacting factors to gene activity. MTA1 participates in physiological functions in the normal and cancer cells. MTA1 is one of the most upregulated proteins in human cancer and associates with cancer progression, aggressive phenotypes, and poor prognosis of cancer patients.

Tripartite motif-containing 24 (TRIM24) also known as transcriptional intermediary factor 1α (TIF1α) is a protein that, in humans, is encoded by the TRIM24 gene.

Sin3A-associated protein, 30kDa, also known as SAP30, is a protein which in humans is encoded by the SAP30 gene.

C-terminal-binding protein 2 also known as CtBP2 is a protein that in humans is encoded by the CTBP2 gene.
Nuclear receptor coregulators are a class of transcription coregulators that have been shown to be involved in any aspect of signaling by any member of the nuclear receptor superfamily. A comprehensive database of coregulators for nuclear receptors and other transcription factors was previously maintained at the Nuclear Receptor Signaling Atlas website which has since been replaced by the Signaling Pathways Project website.
References
- ↑ Glass CK, Rosenfeld MG (2000). "The coregulator exchange in transcriptional functions of nuclear receptors". Genes Dev. 14 (2): 121–41. doi: 10.1101/gad.14.2.121 . PMID 10652267. S2CID 12793980.
- ↑ Schaefer U, Schmeier S, Bajic VB (Jan 2011). "TcoF-DB: dragon database for human transcription co-factors and transcription factor interacting proteins". Nucleic Acids Res. 39 (Database issue): D106-10. doi:10.1093/nar/gkq945. PMC 3013796 . PMID 20965969.
- ↑ Kingston RE, Narlikar GJ (1999). "ATP-dependent remodeling and acetylation as regulators of chromatin fluidity". Genes Dev. 13 (18): 2339–52. doi: 10.1101/gad.13.18.2339 . PMID 10500090.
- 1 2 Choi YB, Ko JK, Shin J (2004). "The transcriptional corepressor, PELP1, recruits HDAC2 and masks histones using two separate domains". J Biol Chem. 279 (49): 50930–41. doi: 10.1074/jbc.M406831200 . PMID 15456770.
- ↑ Shiau AK, Barstad D, Loria PM, Cheng L, Kushner PJ, Agard DA, Greene GL (1998). "The structural basis of estrogen receptor/coactivator recognition and the antagonism of this interaction by tamoxifen". Cell. 95 (7): 927–37. doi: 10.1016/S0092-8674(00)81717-1 . PMID 9875847. S2CID 10265320.
- ↑ Vadlamudi RK, Wang RA, Mazumdar A, Kim Y, Shin J, Sahin A, Kumar R (2001). "Molecular cloning and characterization of PELP1, a novel human coregulator of estrogen receptor alpha". J Biol Chem. 276 (41): 38272–9. doi:10.1074/jbc.M103783200. PMID 11481323.
- ↑ Xu HE, Stanley TB, Montana VG, Lambert MH, Shearer BG, Cobb JE, McKee DD, Galardi CM, Plunket KD, Nolte RT, Parks DJ, Moore JT, Kliewer SA, Willson TM, Stimmel JB (2002). "Structural basis for antagonist-mediated recruitment of nuclear co-repressors by PPARalpha". Nature. 415 (6873): 813–7. Bibcode:2002Natur.415..813X. doi:10.1038/415813a. PMID 11845213. S2CID 4402122.