Heat shock proteins (HSP) are a family of proteins produced by cells in response to exposure to stressful conditions. They were first described in relation to heat shock, but are now known to also be expressed during other stresses including exposure to cold, UV light and during wound healing or tissue remodeling. Many members of this group perform chaperone functions by stabilizing new proteins to ensure correct folding or by helping to refold proteins that were damaged by the cell stress. This increase in expression is transcriptionally regulated. The dramatic upregulation of the heat shock proteins is a key part of the heat shock response and is induced primarily by heat shock factor (HSF). HSPs are found in virtually all living organisms, from bacteria to humans.
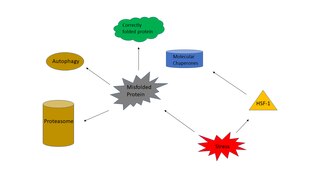
The heat shock response (HSR) is a cell stress response that increases the number of molecular chaperones to combat the negative effects on proteins caused by stressors such as increased temperatures, oxidative stress, and heavy metals. In a normal cell, proteostasis must be maintained because proteins are the main functional units of the cell. Many proteins take on a defined configuration in a process known as protein folding in order to perform their biological functions. If these structures are altered, critical processes could be affected, leading to cell damage or death. The heat shock response can be employed under stress to induce the expression of heat shock proteins (HSP), many of which are molecular chaperones, that help prevent or reverse protein misfolding and provide an environment for proper folding.

The aryl hydrocarbon receptor is a protein that in humans is encoded by the AHR gene. The aryl hydrocarbon receptor is a transcription factor that regulates gene expression. It was originally thought to function primarily as a sensor of xenobiotic chemicals and also as the regulator of enzymes such as cytochrome P450s that metabolize these chemicals. The most notable of these xenobiotic chemicals are aromatic (aryl) hydrocarbons from which the receptor derives its name.
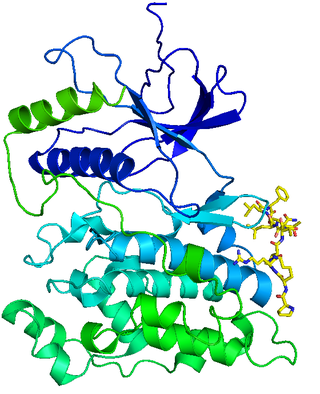
c-Jun N-terminal kinases (JNKs), were originally identified as kinases that bind and phosphorylate c-Jun on Ser-63 and Ser-73 within its transcriptional activation domain. They belong to the mitogen-activated protein kinase family, and are responsive to stress stimuli, such as cytokines, ultraviolet irradiation, heat shock, and osmotic shock. They also play a role in T cell differentiation and the cellular apoptosis pathway. Activation occurs through a dual phosphorylation of threonine (Thr) and tyrosine (Tyr) residues within a Thr-Pro-Tyr motif located in kinase subdomain VIII. Activation is carried out by two MAP kinase kinases, MKK4 and MKK7, and JNK can be inactivated by Ser/Thr and Tyr protein phosphatases. It has been suggested that this signaling pathway contributes to inflammatory responses in mammals and insects.
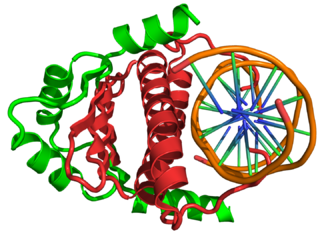
In the field of molecular biology, myocyte enhancer factor-2 (Mef2) proteins are a family of transcription factors which through control of gene expression are important regulators of cellular differentiation and consequently play a critical role in embryonic development. In adult organisms, Mef2 proteins mediate the stress response in some tissues. Mef2 proteins contain both MADS-box and Mef2 DNA-binding domains.

Heat shock 70 kDa protein 1, also termed Hsp72, is a protein that in humans is encoded by the HSPA1A gene. As a member of the heat shock protein 70 family and a chaperone protein, it facilitates the proper folding of newly translated and misfolded proteins, as well as stabilize or degrade mutant proteins. In addition, Hsp72 also facilitates DNA repair. Its functions contribute to biological processes including signal transduction, apoptosis, protein homeostasis, and cell growth and differentiation. It has been associated with an extensive number of cancers, neurodegenerative diseases, cell senescence and aging, and inflammatory diseases such as Diabetes mellitus type 2 and rheumatoid arthritis.

Heat shock protein HSP 90-alpha is a protein that in humans is encoded by the HSP90AA1 gene.

Heat shock factor 1 (HSF1) is a protein that in humans is encoded by the HSF1 gene. HSF1 is highly conserved in eukaryotes and is the primary mediator of transcriptional responses to proteotoxic stress with important roles in non-stress regulation such as development and metabolism.

X-box binding protein 1, also known as XBP1, is a protein which in humans is encoded by the XBP1 gene. The XBP1 gene is located on chromosome 22 while a closely related pseudogene has been identified and localized to chromosome 5. The XBP1 protein is a transcription factor that regulates the expression of genes important to the proper functioning of the immune system and in the cellular stress response.

Activating transcription factor 4 , also known as ATF4, is a protein that in humans is encoded by the ATF4 gene.

Activating transcription factor 6, also known as ATF6, is a protein that, in humans, is encoded by the ATF6 gene and is involved in the unfolded protein response.

Activating transcription factor 2, also known as ATF2, is a protein that, in humans, is encoded by the ATF2 gene.

Heat shock 70 kDa protein 4 is a protein that in humans is encoded by the HSPA4 gene.

DnaJ homolog subfamily B member 1 is a protein that in humans is encoded by the DNAJB1 gene.

Heat shock factor protein 2 is a protein that in humans is encoded by the HSF2 gene.

Cyclic AMP-responsive element-binding protein 3 is a protein that in humans is encoded by the CREB3 gene.

Heat shock factor protein 4 is a protein that in humans is encoded by the HSF4 gene.
The mitochondrial unfolded protein response (UPRmt) is a cellular stress response related to the mitochondria. The UPRmt results from unfolded or misfolded proteins in mitochondria beyond the capacity of chaperone proteins to handle them. The UPRmt can occur either in the mitochondrial matrix or in the mitochondrial inner membrane. In the UPRmt, the mitochondrion will either upregulate chaperone proteins or invoke proteases to degrade proteins that fail to fold properly. UPRmt causes the sirtuin SIRT3 to activate antioxidant enzymes and mitophagy.
CRISPR activation (CRISPRa) is a type of CRISPR tool that uses modified versions of CRISPR effectors without endonuclease activity, with added transcriptional activators on dCas9 or the guide RNAs (gRNAs).
Thermotolerance is the ability of an organism to survive high temperatures. An organism's natural tolerance of heat is their basal thermotolerance. Meanwhile, acquired thermotolerance is defined as an enhanced level of thermotolerance after exposure to a heat stress.















