In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part of biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential.

The origin of replication is a particular sequence in a genome at which replication is initiated. Propagation of the genetic material between generations requires timely and accurate duplication of DNA by semiconservative replication prior to cell division to ensure each daughter cell receives the full complement of chromosomes. This can either involve the replication of DNA in living organisms such as prokaryotes and eukaryotes, or that of DNA or RNA in viruses, such as double-stranded RNA viruses. Synthesis of daughter strands starts at discrete sites, termed replication origins, and proceeds in a bidirectional manner until all genomic DNA is replicated. Despite the fundamental nature of these events, organisms have evolved surprisingly divergent strategies that control replication onset. Although the specific replication origin organization structure and recognition varies from species to species, some common characteristics are shared.
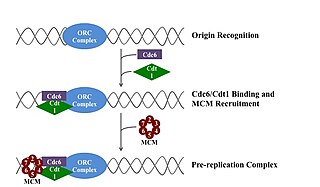
A pre-replication complex (pre-RC) is a protein complex that forms at the origin of replication during the initiation step of DNA replication. Formation of the pre-RC is required for DNA replication to occur. Complete and faithful replication of the genome ensures that each daughter cell will carry the same genetic information as the parent cell. Accordingly, formation of the pre-RC is a very important part of the cell cycle.

S phase (Synthesis phase) is the phase of the cell cycle in which DNA is replicated, occurring between G1 phase and G2 phase. Since accurate duplication of the genome is critical to successful cell division, the processes that occur during S-phase are tightly regulated and widely conserved.
E2F is a group of genes that encodes a family of transcription factors (TF) in higher eukaryotes. Three of them are activators: E2F1, 2 and E2F3a. Six others act as repressors: E2F3b, E2F4-8. All of them are involved in the cell cycle regulation and synthesis of DNA in mammalian cells. E2Fs as TFs bind to the TTTCCCGC consensus binding site in the target promoter sequence.

DNA replication licensing factor MCM6 is a protein that in humans is encoded by the MCM6 gene. MCM6 is one of the highly conserved mini-chromosome maintenance proteins (MCM) that are essential for the initiation of eukaryotic genome replication.
A licensing factor is a protein or complex of proteins that allows an origin of replication to begin DNA replication at that site. Licensing factors primarily occur in eukaryotic cells, since bacteria use simpler systems to initiate replication. However, many archaea use homologues of eukaryotic licensing factors to initiate replication.

Geminin, DNA replication inhibitor, also known as GMNN, is a protein in humans encoded by the GMNN gene. A nuclear protein present in most eukaryotes and highly conserved across species, numerous functions have been elucidated for geminin including roles in metazoan cell cycle, cellular proliferation, cell lineage commitment, and neural differentiation. One example of its function is the inhibition of Cdt1.
In molecular biology, origin recognition complex (ORC) is a multi-subunit DNA binding complex that binds in all eukaryotes and archaea in an ATP-dependent manner to origins of replication. The subunits of this complex are encoded by the ORC1, ORC2, ORC3, ORC4, ORC5 and ORC6 genes. ORC is a central component for eukaryotic DNA replication, and remains bound to chromatin at replication origins throughout the cell cycle.

Cyclin-dependent kinase 2, also known as cell division protein kinase 2, or Cdk2, is an enzyme that in humans is encoded by the CDK2 gene. The protein encoded by this gene is a member of the cyclin-dependent kinase family of Ser/Thr protein kinases. This protein kinase is highly similar to the gene products of S. cerevisiae cdc28, and S. pombe cdc2, also known as Cdk1 in humans. It is a catalytic subunit of the cyclin-dependent kinase complex, whose activity is restricted to the G1-S phase of the cell cycle, where cells make proteins necessary for mitosis and replicate their DNA. This protein associates with and is regulated by the regulatory subunits of the complex including cyclin E or A. Cyclin E binds G1 phase Cdk2, which is required for the transition from G1 to S phase while binding with Cyclin A is required to progress through the S phase. Its activity is also regulated by phosphorylation. Multiple alternatively spliced variants and multiple transcription initiation sites of this gene have been reported. The role of this protein in G1-S transition has been recently questioned as cells lacking Cdk2 are reported to have no problem during this transition.

Eukaryotic DNA replication is a conserved mechanism that restricts DNA replication to once per cell cycle. Eukaryotic DNA replication of chromosomal DNA is central for the duplication of a cell and is necessary for the maintenance of the eukaryotic genome.

The minichromosome maintenance protein complex (MCM) is a DNA helicase essential for genomic DNA replication. Eukaryotic MCM consists of six gene products, Mcm2–7, which form a heterohexamer. As a critical protein for cell division, MCM is also the target of various checkpoint pathways, such as the S-phase entry and S-phase arrest checkpoints. Both the loading and activation of MCM helicase are strictly regulated and are coupled to cell growth cycles. Deregulation of MCM function has been linked to genomic instability and a variety of carcinomas.

DNA replication licensing factor MCM7 is a protein that in humans is encoded by the MCM7 gene.

DNA replication licensing factor MCM2 is a protein that in humans is encoded by the MCM2 gene.

DNA replication licensing factor MCM4 is a protein that in humans is encoded by the MCM4 gene.

CDT1 is a protein that in humans is encoded by the CDT1 gene. It is a licensing factor that functions to limit DNA from replicating more than once per cell cycle.

Origin recognition complex subunit 2 is a protein that is encoded by the ORC2 (ORC2L) gene in humans.

Cell division cycle 7-related protein kinase is an enzyme that in humans is encoded by the CDC7 gene. The Cdc7 kinase is involved in regulation of the cell cycle at the point of chromosomal DNA replication. The gene CDC7 appears to be conserved throughout eukaryotic evolution; this means that most eukaryotic cells have the Cdc7 kinase protein.

In cell biology, eukaryotes possess a regulatory system that ensures that DNA replication occurs only once per cell cycle.

DNA re-replication is an undesirable and possibly fatal occurrence in eukaryotic cells in which the genome is replicated more than once per cell cycle. Rereplication is believed to lead to genomic instability and has been implicated in the pathologies of a variety of human cancers. To prevent rereplication, eukaryotic cells have evolved multiple, overlapping mechanisms to inhibit chromosomal DNA from being partially or fully rereplicated in a given cell cycle. These control mechanisms rely on cyclin-dependent kinase (CDK) activity. DNA replication control mechanisms cooperate to prevent the relicensing of replication origins and to activate cell cycle and DNA damage checkpoints. DNA rereplication must be strictly regulated to ensure that genomic information is faithfully transmitted through successive generations.