
An electron transport chain (ETC) is a series of protein complexes and other molecules that transfer electrons from electron donors to electron acceptors via redox reactions (both reduction and oxidation occurring simultaneously) and couples this electron transfer with the transfer of protons (H+ ions) across a membrane. The electrons that are transferred from NADH and FADH2 to the ETC involves four multi-subunit large enzymes complexes and two mobile electron carriers. Many of the enzymes in the electron transport chain are embedded within the membrane.

Ribonucleotide reductase (RNR), also known as ribonucleoside diphosphate reductase (rNDP), is an enzyme that catalyzes the formation of deoxyribonucleotides from ribonucleotides. It catalyzes this formation by removing the 2'-hydroxyl group of the ribose ring of nucleoside diphosphates. This reduction produces deoxyribonucleotides. Deoxyribonucleotides in turn are used in the synthesis of DNA. The reaction catalyzed by RNR is strictly conserved in all living organisms. Furthermore, RNR plays a critical role in regulating the total rate of DNA synthesis so that DNA to cell mass is maintained at a constant ratio during cell division and DNA repair. A somewhat unusual feature of the RNR enzyme is that it catalyzes a reaction that proceeds via a free radical mechanism of action. The substrates for RNR are ADP, GDP, CDP and UDP. dTDP is synthesized by another enzyme from dTMP.
Ferredoxins are iron–sulfur proteins that mediate electron transfer in a range of metabolic reactions. The term "ferredoxin" was coined by D.C. Wharton of the DuPont Co. and applied to the "iron protein" first purified in 1962 by Mortenson, Valentine, and Carnahan from the anaerobic bacterium Clostridium pasteurianum.
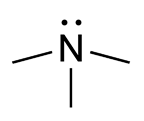
Trimethylaminuria (TMAU), also known as fish odor syndrome or fish malodor syndrome, is a rare metabolic disorder that causes a defect in the normal production of an enzyme named flavin-containing monooxygenase 3 (FMO3). When FMO3 is not working correctly or if not enough enzyme is produced, the body loses the ability to properly convert trimethylamine (TMA) from precursor compounds in food digestion into trimethylamine oxide (TMAO), through a process called N-oxidation. Trimethylamine then builds up and is released in the person's sweat, urine, and breath, giving off a fishy odor. Primary trimethylaminuria is caused by genetic mutations that affect the FMO3 function of the liver. Symptoms matching TMAU can also occur when there is no genetic cause, yet excessive TMA is excreted - this has been described as secondary trimethylaminuria (TMAU2). TMAU2 can be caused by a precursor overload, hormonal issues related to menstrual cycles, liver damage, or liver and kidney failure. As a symptom rather than a disease, TMAU2 is temporary and will resolve as the underlying cause is remedied.
DMSO reductase is a molybdenum-containing enzyme that catalyzes reduction of dimethyl sulfoxide (DMSO) to dimethyl sulfide (DMS). This enzyme serves as the terminal reductase under anaerobic conditions in some bacteria, with DMSO being the terminal electron acceptor. During the course of the reaction, the oxygen atom in DMSO is transferred to molybdenum, and then reduced to water.

Sulfite oxidase is an enzyme in the mitochondria of all eukaryotes, with exception of the yeasts. It oxidizes sulfite to sulfate and, via cytochrome c, transfers the electrons produced to the electron transport chain, allowing generation of ATP in oxidative phosphorylation. This is the last step in the metabolism of sulfur-containing compounds and the sulfate is excreted.
Microbial metabolism is the means by which a microbe obtains the energy and nutrients it needs to live and reproduce. Microbes use many different types of metabolic strategies and species can often be differentiated from each other based on metabolic characteristics. The specific metabolic properties of a microbe are the major factors in determining that microbe's ecological niche, and often allow for that microbe to be useful in industrial processes or responsible for biogeochemical cycles.

Nitrate reductases are molybdoenzymes that reduce nitrate to nitrite. This reaction is critical for the production of protein in most crop plants, as nitrate is the predominant source of nitrogen in fertilized soils.

Trimethylamine N-oxide (TMAO) is an organic compound with the formula (CH3)3NO. It is in the class of amine oxides. Although the anhydrous compound is known, trimethylamine N-oxide is usually encountered as the dihydrate. Both the anhydrous and hydrated materials are white, water-soluble solids.

2,4 Dienoyl-CoA reductase also known as DECR1 is an enzyme which in humans is encoded by the DECR1 gene which resides on chromosome 8. This enzyme catalyzes the following reactions

Flavin-containing monooxygenase 3 (FMO3), also known as dimethylaniline monooxygenase [N-oxide-forming] 3 and trimethylamine monooxygenase, is a flavoprotein enzyme (EC 1.14.13.148) that in humans is encoded by the FMO3 gene. This enzyme catalyzes the following chemical reaction, among others:
Formate dehydrogenases are a set of enzymes that catalyse the oxidation of formate to carbon dioxide, donating the electrons to a second substrate, such as NAD+ in formate:NAD+ oxidoreductase (EC 1.17.1.9) or to a cytochrome in formate:ferricytochrome-b1 oxidoreductase (EC 1.2.2.1). This family of enzymes has attracted attention as inspiration or guidance on methods for the carbon dioxide fixation, relevant to global warming.
In enzymology, an ethylbenzene hydroxylase (EC 1.17.99.2) is an enzyme that catalyzes the chemical reaction
Nitric oxide reductase, an enzyme, catalyzes the reduction of nitric oxide (NO) to nitrous oxide (N2O). The enzyme participates in nitrogen metabolism and in the microbial defense against nitric oxide toxicity. The catalyzed reaction may be dependent on different participating small molecules: Cytochrome c (EC: 1.7.2.5, Nitric oxide reductase (cytochrome c)), NADPH (EC:1.7.1.14), or Menaquinone (EC:1.7.5.2).

Cytochrome c nitrite reductase (ccNiR) is a bacterial enzyme that catalyzes the six electron reduction of nitrite to ammonia; an important step in the biological nitrogen cycle. The enzyme catalyses the second step in the two step conversion of nitrate to ammonia, which allows certain bacteria to use nitrite as a terminal electron acceptor, rather than oxygen, during anaerobic conditions. During this process, ccNiR draws electrons from the quinol pool, which are ultimately provided by a dehydrogenase such as formate dehydrogenase or hydrogenase. These dehydrogenases are responsible for generating a proton motive force.
In enzymology, a nitrite reductase (NO-forming) (EC 1.7.2.1) is an enzyme that catalyzes the chemical reaction
In enzymology, a trimethylamine-N-oxide reductase (cytochrome c) (EC 1.7.2.3) is an enzyme that catalyzes the chemical reaction
Nitrate reductase (quinone) (EC 1.7.5.1, nitrate reductase A, nitrate reductase Z, quinol/nitrate oxidoreductase, quinol-nitrate oxidoreductase, quinol:nitrate oxidoreductase, NarA, NarZ, NarGHI) is an enzyme with systematic name nitrite:quinone oxidoreductase. This enzyme catalyses the following chemical reaction
Molybdenum cofactor cytidylyltransferase is an enzyme with systematic name CTP:molybdenum cofactor cytidylyltransferase. This enzyme catalyses the following chemical reaction:
The Disulfide bond oxidoreductase D (DsbD) family is a member of the Lysine Exporter (LysE) Superfamily. A representative list of proteins belonging to the DsbD family can be found in the Transporter Classification Base.











