Related Research Articles
An abzyme, also called catmab, and most often called catalytic antibody or sometimes catab, is a monoclonal antibody with catalytic activity. Abzymes are usually raised in lab animals immunized against synthetic haptens, but some natural abzymes can be found in normal humans and in patients with autoimmune diseases such as systemic lupus erythematosus, where they can bind to and hydrolyze DNA. To date abzymes display only weak, modest catalytic activity and have not proved to be of any practical use. They are, however, subjects of considerable academic interest. Studying them has yielded important insights into reaction mechanisms, enzyme structure and function, catalysis, and the immune system itself.
Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine at the P1 position.

A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism, and cell signaling.

In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate, the binding site, and residues that catalyse a reaction of that substrate, the catalytic site. Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes.
Serine proteases are enzymes that cleave peptide bonds in proteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like.
A metalloproteinase, or metalloprotease, is any protease enzyme whose catalytic mechanism involves a metal. An example is ADAM12 which plays a significant role in the fusion of muscle cells during embryo development, in a process known as myogenesis.

Papain, also known as papaya proteinase I, is a cysteine protease enzyme present in papaya and mountain papaya. It is the namesake member of the papain-like protease family.
Enteropeptidase is an enzyme produced by cells of the duodenum and is involved in digestion in humans and other animals. Enteropeptidase converts trypsinogen into its active form trypsin, resulting in the subsequent activation of pancreatic digestive enzymes. Absence of enteropeptidase results in intestinal digestion impairment.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.
Cysteine proteases, also known as thiol proteases, are hydrolase enzymes that degrade proteins. These proteases share a common catalytic mechanism that involves a nucleophilic cysteine thiol in a catalytic triad or dyad.

An oxyanion hole is a pocket in the active site of an enzyme that stabilizes transition state negative charge on a deprotonated oxygen or alkoxide. The pocket typically consists of backbone amides or positively charged residues. Stabilising the transition state lowers the activation energy necessary for the reaction, and so promotes catalysis. For example, proteases such as chymotrypsin contain an oxyanion hole to stabilise the tetrahedral intermediate anion formed during proteolysis and protects substrate's negatively charged oxygen from water molecules. Additionally, it may allow for insertion or positioning of a substrate, which would suffer from steric hindrance if it could not occupy the hole. Enzymes that catalyse multi-step reactions can have multiple oxyanion holes that stabilise different transition states in the reaction.

Subtilisin is a protease initially obtained from Bacillus subtilis.
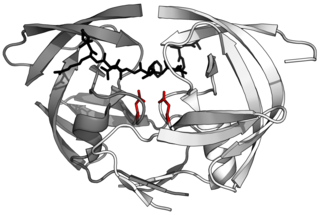
Aspartic proteases are a catalytic type of protease enzymes that use an activated water molecule bound to one or more aspartate residues for catalysis of their peptide substrates. In general, they have two highly conserved aspartates in the active site and are optimally active at acidic pH. Nearly all known aspartyl proteases are inhibited by pepstatin.

HIV-1 protease (PR) is a retroviral aspartyl protease (retropepsin), an enzyme involved with peptide bond hydrolysis in retroviruses, that is essential for the life-cycle of HIV, the retrovirus that causes AIDS. HIV protease cleaves newly synthesized polyproteins at nine cleavage sites to create the mature protein components of an HIV virion, the infectious form of a virus outside of the host cell. Without effective HIV protease, HIV virions remain uninfectious.

Threonine proteases are a family of proteolytic enzymes harbouring a threonine (Thr) residue within the active site. The prototype members of this class of enzymes are the catalytic subunits of the proteasome, however the acyltransferases convergently evolved the same active site geometry and mechanism.
C-terminal processing peptidase is an enzyme. This enzyme catalyses the following chemical reaction
Mannan-binding lectin-associated serine protease-2 is an enzyme. This enzyme catalyses the following chemical reaction

The 3C-like protease (3CLpro) or main protease (Mpro), formally known as C30 endopeptidase or 3-chymotrypsin-like protease, is the main protease found in coronaviruses. It cleaves the coronavirus polyprotein at eleven conserved sites. It is a cysteine protease and a member of the PA clan of proteases. It has a cysteine-histidine catalytic dyad at its active site and cleaves a Gln–(Ser/Ala/Gly) peptide bond.

Aspergilloglutamic peptidase, also called aspergillopepsin II is a proteolytic enzyme. The enzyme was previously thought be an aspartic protease, but it was later shown to be a glutamic protease with a catalytic Glu residue at the active site, and was therefore renamed aspergilloglutamic peptidase.

The PA clan is the largest group of proteases with common ancestry as identified by structural homology. Members have a chymotrypsin-like fold and similar proteolysis mechanisms but can have identity of <10%. The clan contains both cysteine and serine proteases. PA clan proteases can be found in plants, animals, fungi, eubacteria, archaea and viruses.
References
- ↑ Chen P, Tsuge H, Almassy RJ, Gribskov CL, Katoh S, Vanderpool DL, Margosiak SA, Pinko C, Matthews DA, Kan CC (September 1996). "Structure of the human cytomegalovirus protease catalytic domain reveals a novel serine protease fold and catalytic triad". Cell. 86 (5): 835–43. doi: 10.1016/s0092-8674(00)80157-9 . PMID 8797829.
- ↑ Darke PL, Rawlings ND, Woessner JF (1998). "Herpesvirus assemblin". In Barrett AJ (ed.). Handbook of Proteolytic Enzymes. London: Academic Press. pp. 470–472.