A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create two identical DNA duplexes from a single original DNA duplex. During this process, DNA polymerase "reads" the existing DNA strands to create two new strands that match the existing ones. These enzymes catalyze the chemical reaction
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA segment called a primer complementary to a ssDNA template. After this elongation, the RNA piece is removed by a 5' to 3' exonuclease and refilled with DNA.

A ρ factor is a prokaryotic protein involved in the termination of transcription. Rho factor binds to the transcription terminator pause site, an exposed region of single stranded RNA after the open reading frame at C-rich/G-poor sequences that lack obvious secondary structure.

A nuclease is an enzyme capable of cleaving the phosphodiester bonds between nucleotides of nucleic acids. Nucleases variously effect single and double stranded breaks in their target molecules. In living organisms, they are essential machinery for many aspects of DNA repair. Defects in certain nucleases can cause genetic instability or immunodeficiency. Nucleases are also extensively used in molecular cloning.
dnaQ is the gene encoding the ε subunit of DNA polymerase III in Escherichia coli. The ε subunit is one of three core proteins in the DNA polymerase complex. It functions as a 3’→5’ DNA directed proofreading exonuclease that removes incorrectly incorporated bases during replication. dnaQ may also be referred to as mutD.

The nucleoid is an irregularly shaped region within the prokaryotic cell that contains all or most of the genetic material. The chromosome of a prokaryote is circular, and its length is very large compared to the cell dimensions needing it to be compacted in order to fit. In contrast to the nucleus of a eukaryotic cell, it is not surrounded by a nuclear membrane. Instead, the nucleoid forms by condensation and functional arrangement with the help of chromosomal architectural proteins and RNA molecules as well as DNA supercoiling. The length of a genome widely varies and a cell may contain multiple copies of it.
DNA gyrase, or simply gyrase, is an enzyme within the class of topoisomerase and is a subclass of Type II topoisomerases that reduces topological strain in an ATP dependent manner while double-stranded DNA is being unwound by elongating RNA-polymerase or by helicase in front of the progressing replication fork. The enzyme causes negative supercoiling of the DNA or relaxes positive supercoils. It does so by looping the template so as to form a crossing, then cutting one of the double helices and passing the other through it before releasing the break, changing the linking number by two in each enzymatic step. This process occurs in bacteria, whose single circular DNA is cut by DNA gyrase and the two ends are then twisted around each other to form supercoils. Gyrase is also found in eukaryotic plastids: it has been found in the apicoplast of the malarial parasite Plasmodium falciparum and in chloroplasts of several plants. Bacterial DNA gyrase is the target of many antibiotics, including nalidixic acid, novobiocin, and ciprofloxacin.
Bacterial translation is the process by which messenger RNA is translated into proteins in bacteria.

Type II topoisomerases are topoisomerases that cut both strands of the DNA helix simultaneously in order to manage DNA tangles and supercoils. They use the hydrolysis of ATP, unlike Type I topoisomerase. In this process, these enzymes change the linking number of circular DNA by ±2.

Prokaryotic DNA Replication is the process by which a prokaryote duplicates its DNA into another copy that is passed on to daughter cells. Although it is often studied in the model organism E. coli, other bacteria show many similarities. Replication is bi-directional and originates at a single origin of replication (OriC). It consists of three steps: Initiation, elongation, and termination.

Replication protein A (RPA) is the major protein that binds to single-stranded DNA (ssDNA) in eukaryotic cells. In vitro, RPA shows a much higher affinity for ssDNA than RNA or double-stranded DNA.

T7 DNA polymerase is an enzyme used during the DNA replication of the T7 bacteriophage. During this process, the DNA polymerase “reads” existing DNA strands and creates two new strands that match the existing ones. The T7 DNA polymerase requires a host factor, E. coli thioredoxin, in order to carry out its function. This helps stabilize the binding of the necessary protein to the primer-template to improve processivity by more than 100-fold, which is a feature unique to this enzyme. It is a member of the Family A DNA polymerases, which include E. coli DNA polymerase I and Taq DNA polymerase.
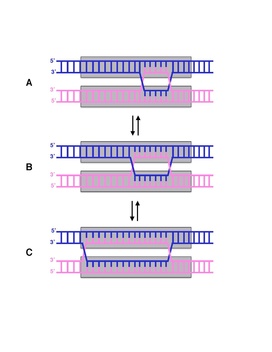
Branch migration is the process by which base pairs on homologous DNA strands are consecutively exchanged at a Holliday junction, moving the branch point up or down the DNA sequence. Branch migration is the second step of genetic recombination, following the exchange of two single strands of DNA between two homologous chromosomes. The process is random, and the branch point can be displaced in either direction on the strand, influencing the degree of which the genetic material is exchanged. Branch migration can also be seen in DNA repair and replication, when filling in gaps in the sequence. It can also be seen when a foreign piece of DNA invades the strand.
The RecF pathway, also called the RecFOR pathway, is a pathway of homologous recombination that repairs DNA in bacteria. It repairs breaks that occur on only one of DNA's two strands, known as single-strand gaps. The RecF pathway can also repair double-strand breaks in DNA when the RecBCD pathway, another pathway of homologous recombination in bacteria, is inactivated by mutations. Like the RecBCD pathway, the RecF pathway requires RecA for strand invasion. The two pathways are also similar in their phases of branch migration, in which the Holliday junction slides in one direction, and resolution, in which the Holliday junctions are cleaved apart by enzymes.

In molecular biology the SeqA protein is found in bacteria and archaea. The function of this protein is highly important in DNA replication. The protein negatively regulates the initiation of DNA replication at the origin of replication, in Escherichia coli, OriC. Additionally the protein plays a further role in sequestration. The importance of this protein is vital, without its help in DNA replication, cell division and other crucial processes could not occur. This protein domain is thought to be part of a much larger protein complex which includes other proteins such as SeqB.
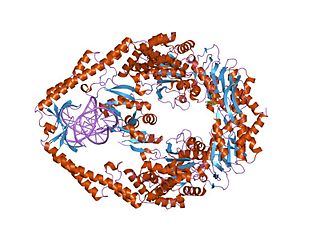
MutS is a mismatch DNA repair protein, originally described in Escherichia coli.
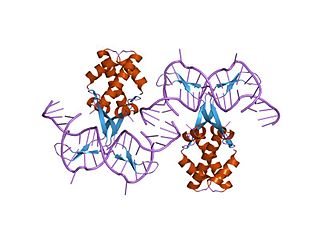
In molecular biology, bacterial DNA binding proteins are a family of small, usually basic proteins of about 90 residues that bind DNA and are known as histone-like proteins. Since bacterial binding proteins have a diversity of functions, it has been difficult to develop a common function for all of them. They are commonly referred to as histone-like and have many similar traits with the eukaryotic histone proteins. Eukaryotic histones package DNA to help it to fit in the nucleus, and they are known to be the most conserved proteins in nature. Examples include the HU protein in Escherichia coli, a dimer of closely related alpha and beta chains and in other bacteria can be a dimer of identical chains. HU-type proteins have been found in a variety of eubacteria and archaebacteria, and are also encoded in the chloroplast genome of some algae. The integration host factor (IHF), a dimer of closely related chains which is suggested to function in genetic recombination as well as in translational and transcriptional control is found in Enterobacteria and viral proteins including the African swine fever virus protein A104R.

In molecular biology, the CRM domain is an approximately 100-amino acid RNA-binding domain. The name CRM has been suggested to reflect the functions established for four characterised members of the family: Zea mays (Maize) CRS1, CAF1 and CAF2 proteins and the Escherichia coli protein YhbY. Proteins containing the CRM domain are found in eubacteria, archaea, and plants. The CRM domain is represented as a stand-alone protein in archaea and bacteria, and in single- and multi-domain proteins in plants. It has been suggested that prokaryotic CRM proteins existed as ribosome-associated proteins prior to the divergence of archaea and bacteria, and that they were co-opted in the plant lineage as RNA binding modules by incorporation into diverse protein contexts. Plant CRM domains are predicted to reside not only in the chloroplast, but also in the mitochondrion and the nucleo/cytoplasmic compartment. The diversity of the CRM domain family in plants suggests a diverse set of RNA targets.
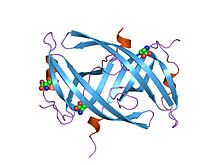
Single-stranded binding proteins (SSBs) are a class of proteins that have been identified in both viruses and organisms from bacteria to humans.

The universal stress protein (USP) domain is a superfamily of conserved genes which can be found in bacteria, archaea, fungi, protozoa and plants. Proteins containing the domain are induced by many environmental stressors such as nutrient starvation, drought, extreme temperatures, high salinity, and the presence of uncouplers, antibiotics and metals.