External links
- Endodeoxyribonucleases at the US National Library of Medicine Medical Subject Headings (MeSH)
Endodeoxyribonuclease are both endonucleases and deoxyribonucleases. They catalyze cleavage of the phosphodiester bonds in DNA. They are classified with EC numbers 3.1.21 through 3.1.25.
Examples include:
A restriction enzyme, restriction endonuclease, or restrictase is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each sugar-phosphate backbone of the DNA double helix.

A deoxyribonuclease is an enzyme that catalyzes the hydrolytic cleavage of phosphodiester linkages in the DNA backbone, thus degrading DNA. Deoxyribonucleases are one type of nuclease, a generic term for enzymes capable of hydrolyzing phosphodiester bonds that link nucleotides. A wide variety of deoxyribonucleases are known, which differ in their substrate specificities, chemical mechanisms, and biological functions.

A nuclease is an enzyme capable of cleaving the phosphodiester bonds between nucleotides of nucleic acids. Nucleases variously affect single and double stranded breaks in their target molecules. In living organisms, they are essential machinery for many aspects of DNA repair. Defects in certain nucleases can cause genetic instability or immunodeficiency. Nucleases are also extensively used in molecular cloning.
Endonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain. Some, such as deoxyribonuclease I, cut DNA relatively nonspecifically, while many, typically called restriction endonucleases or restriction enzymes, cleave only at very specific nucleotide sequences. Endonucleases differ from exonucleases, which cleave the ends of recognition sequences instead of the middle (endo) portion. Some enzymes known as "exo-endonucleases", however, are not limited to either nuclease function, displaying qualities that are both endo- and exo-like. Evidence suggests that endonuclease activity experiences a lag compared to exonuclease activity.

Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease.
An esterase is a hydrolase enzyme that splits esters into an acid and an alcohol in a chemical reaction with water called hydrolysis.
UvrABC endonuclease is a multienzyme complex in bacteria involved in DNA repair by nucleotide excision repair, and it is, therefore, sometimes called an excinuclease. This UvrABC repair process, sometimes called the short-patch process, involves the removal of twelve nucleotides where a genetic mutation has occurred followed by a DNA polymerase, replacing these aberrant nucleotides with the correct nucleotides and completing the DNA repair. The subunits for this enzyme are encoded in the uvrA, uvrB, and uvrC genes. This enzyme complex is able to repair many different types of damage, including cyclobutyl dimer formation.
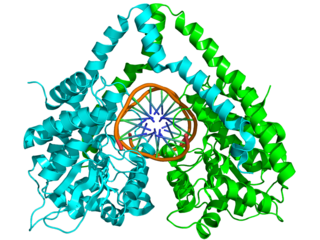
HindIII (pronounced "Hin D Three") is a type II site-specific deoxyribonuclease restriction enzyme isolated from Haemophilus influenzae that cleaves the DNA palindromic sequence AAGCTT in the presence of the cofactor Mg2+ via hydrolysis.
Flap endonucleases are a class of nucleolytic enzymes that act as both 5'-3' exonucleases and structure-specific endonucleases on specialised DNA structures that occur during the biological processes of DNA replication, DNA repair, and DNA recombination. Flap endonucleases have been identified in eukaryotes, prokaryotes, archaea, and some viruses. Organisms can have more than one FEN homologue; this redundancy may give an indication of the importance of these enzymes. In prokaryotes, the FEN enzyme is found as an N-terminal domain of DNA polymerase I, but some prokaryotes appear to encode a second homologue.

Nuclease S1 is an endonuclease enzyme that splits single-stranded DNA (ssDNA) and RNA into oligo- or mononucleotides. This enzyme catalyses the following chemical reaction
Deoxyribonuclease IV (phage-T4-induced) is a kind of Endonuclease that catalyzes the degradation nucleotides in DsDNA by attacking the 5'-terminal end.
In enzymology, DNA-(apurinic or apyrimidinic site) lyase, also referred to as DNA-(apurinic or apyrimidinic site) 5'-phosphomonoester-lyase or DNA AP lyase is a class of enzyme that catalyzes the chemical reaction of the cleavage of the C3'-O-P bond 3' from the apurinic or apyrimidinic site in DNA via beta-elimination reaction, leaving a 3'-terminal unsaturated sugar and a product with a terminal 5'-phosphate. In the 1970s, this class of enzyme was found to repair at apurinic or apyrimidinic DNA sites in E. coli and in mammalian cells. The major active enzyme of this class in bacteria, and specifically, E. coli is endonuclease type III. This enzyme is part of a family of lyases that cleave carbon-oxygen bonds.
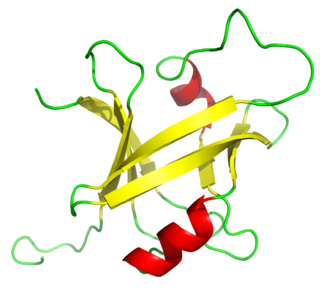
The B3 DNA binding domain (DBD) is a highly conserved domain found exclusively in transcription factors combined with other domains. It consists of 100-120 residues, includes seven beta strands and two alpha helices that form a DNA-binding pseudobarrel protein fold ; it interacts with the major groove of DNA.
Meganucleases are endodeoxyribonucleases characterized by a large recognition site ; as a result this site generally occurs only once in any given genome. For example, the 18-base pair sequence recognized by the I-SceI meganuclease would on average require a genome twenty times the size of the human genome to be found once by chance. Meganucleases are therefore considered to be the most specific naturally occurring restriction enzymes.
DNA ends refer to the properties of the end of DNA molecules, which may be sticky or blunt based on the enzyme which cuts the DNA. The restriction enzyme belong to a larger class of enzymes called exonucleases and endonucleases. Exonucleases remove nucleotide from ends whereas endonuclease cuts at specific position within the DNA.
CC-preferring endodeoxyribonuclease is an enzyme. This enzyme catalyses the following chemical reaction
Crossover junction endodeoxyribonuclease, also known as Holliday junction resolvase, Holliday junction endonuclease, Holliday junction-cleaving endonuclease, Holliday junction-resolving endoribonuclease, crossover junction endoribonuclease, and cruciform-cutting endonuclease, is an enzyme involved in DNA repair and homologous recombination. Specifically, it performs endonucleolytic cleavage that results in single-stranded crossover between two homologous DNA molecules at the Holliday junction to produce recombinant DNA products for chromosomal segregation. This process is known as Holliday junction resolution.
Deoxyribonuclease X is an enzyme. This enzyme catalyses the following chemical reaction
Deoxyribonuclease is an enzyme. This enzyme catalyses the following chemical reaction

Monodnaviria is a realm of viruses that includes all single-stranded DNA viruses that encode an endonuclease of the HUH superfamily that initiates rolling circle replication of the circular viral genome. Viruses descended from such viruses are also included in the realm, including certain linear single-stranded DNA (ssDNA) viruses and circular double-stranded DNA (dsDNA) viruses. These atypical members typically replicate through means other than rolling circle replication.