In biochemistry, isomerases are a general class of enzymes that convert a molecule from one isomer to another. Isomerases facilitate intramolecular rearrangements in which bonds are broken and formed. The general form of such a reaction is as follows:
In organic chemistry, a tetrose is a monosaccharide with 4 carbon atoms. They have either an aldehyde functional group in position 1 (aldotetroses) or a ketone group in position 2 (ketotetroses).
Enoyl-CoA-(∆) isomerase (EC 5.3.3.8, also known as dodecenoyl-CoA- isomerase, 3,2-trans-enoyl-CoA isomerase, ∆3 ,∆2 -enoyl-CoA isomerase, or acetylene-allene isomerase, is an enzyme that catalyzes the conversion of cis- or trans-double bonds of coenzyme A bound fatty acids at gamma-carbon to trans double bonds at beta-carbon as below:
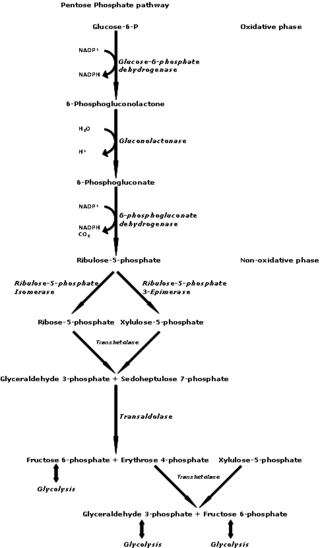
The pentose phosphate pathway is a metabolic pathway parallel to glycolysis. It generates NADPH and pentoses as well as ribose 5-phosphate, a precursor for the synthesis of nucleotides. While the pentose phosphate pathway does involve oxidation of glucose, its primary role is anabolic rather than catabolic. The pathway is especially important in red blood cells (erythrocytes). The reactions of the pathway were elucidated in the early 1950s by Bernard Horecker and co-workers.

Triose-phosphate isomerase is an enzyme that catalyzes the reversible interconversion of the triose phosphate isomers dihydroxyacetone phosphate and D-glyceraldehyde 3-phosphate.

Glucose-6-phosphate dehydrogenase (G6PD or G6PDH) (EC 1.1.1.49) is a cytosolic enzyme that catalyzes the chemical reaction

Amino acid biosynthesis is the set of biochemical processes by which the amino acids are produced. The substrates for these processes are various compounds in the organism's diet or growth media. Not all organisms are able to synthesize all amino acids. For example, humans can synthesize 11 of the 20 standard amino acids. These 11 are called the non-essential amino acids.

In enzymology, aldose reductase is an enzyme in humans encoded by the gene AKR1B1. It is an cytosolic NADPH-dependent oxidoreductase that catalyzes the reduction of a variety of aldehydes and carbonyls, including monosaccharides, and primarily known for catalyzing the reduction of glucose to sorbitol, the first step in polyol pathway of glucose metabolism.

6-Phosphogluconate dehydrogenase (6PGD) is an enzyme in the pentose phosphate pathway. It forms ribulose 5-phosphate from 6-phosphogluconate:

Transaldolase is an enzyme of the non-oxidative phase of the pentose phosphate pathway. In humans, transaldolase is encoded by the TALDO1 gene.

Ribose 5-phosphate (R5P) is both a product and an intermediate of the pentose phosphate pathway. The last step of the oxidative reactions in the pentose phosphate pathway is the production of ribulose 5-phosphate. Depending on the body's state, ribulose 5-phosphate can reversibly isomerize to ribose 5-phosphate. Ribulose 5-phosphate can alternatively undergo a series of isomerizations as well as transaldolations and transketolations that result in the production of other pentose phosphates as well as fructose 6-phosphate and glyceraldehyde 3-phosphate.

6-Phosphogluconolactone is an intermediate in the pentose phosphate pathway (PPP).

Phosphopentose epimerase encoded in humans by the RPE gene is a metalloprotein that catalyzes the interconversion between D-ribulose 5-phosphate and D-xylulose 5-phosphate.

Sucrose phosphorylase is an important enzyme in the metabolism of sucrose and regulation of other metabolic intermediates. Sucrose phosphorylase is in the class of hexosyltransferases. More specifically it has been placed in the retaining glycoside hydrolases family although it catalyzes a transglycosidation rather than hydrolysis. Sucrose phosphorylase catalyzes the conversion of sucrose to D-fructose and α-D-glucose-1-phosphate. It has been shown in multiple experiments that the enzyme catalyzes this conversion by a double displacement mechanism.
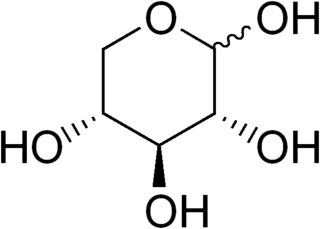
D-Xylose is a five-carbon aldose that can be catabolized or metabolized into useful products by a variety of organisms.

In enzymology, a D-xylulose reductase (EC 1.1.1.9) is an enzyme that is classified as an Oxidoreductase (EC 1) specifically acting on the CH-OH group of donors (EC 1.1.1) that uses NAD+ or NADP+ as an acceptor (EC 1.1.1.9). This enzyme participates in pentose and glucuronate interconversions; a set of metabolic pathways that involve converting pentose sugars and glucuronate into other compounds.
In enzymology, a phosphogluconate dehydrogenase (decarboxylating) (EC 1.1.1.44) is an enzyme that catalyzes the chemical reaction

Ribose-5-phosphate isomerase (Rpi) encoded by the RPIA gene is an enzyme that catalyzes the conversion between ribose-5-phosphate (R5P) and ribulose-5-phosphate (Ru5P). It is a member of a larger class of isomerases which catalyze the interconversion of chemical isomers. It plays a vital role in biochemical metabolism in both the pentose phosphate pathway and the Calvin cycle. The systematic name of this enzyme class is D-ribose-5-phosphate aldose-ketose-isomerase.
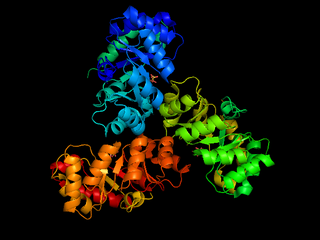
The enzyme 2-dehydro-3-deoxy-phosphogluconate aldolase, commonly known as KDPG aldolase, catalyzes the chemical reaction

In enzymology, a xylose isomerase is an enzyme that catalyzes the interconversion of D-xylose and D-xylulose. This enzyme belongs to the family of isomerases, specifically those intramolecular oxidoreductases interconverting aldoses and ketoses. The isomerase has now been observed in nearly a hundred species of bacteria. Xylose-isomerases are also commonly called glucose isomerase or fructose isomerases due to their ability to interconvert glucose and fructose. The systematic name of this enzyme class is α-D-xylopyranose aldose-ketose-isomerase. Other names in common use include D-xylose isomerase, D-xylose ketoisomerase, and D-xylose ketol-isomerase.



















