
Proteasomes are protein complexes which degrade unneeded or damaged proteins by proteolysis, a chemical reaction that breaks peptide bonds. Enzymes that help such reactions are called proteases.

In molecular biology, molecular chaperones are proteins that assist the conformational folding or unfolding of large proteins or macromolecular protein complexes. There are a number of classes of molecular chaperones, all of which function to assist large proteins in proper protein folding during or after synthesis, and after partial denaturation. Chaperones are also involved in the translocation of proteins for proteolysis.
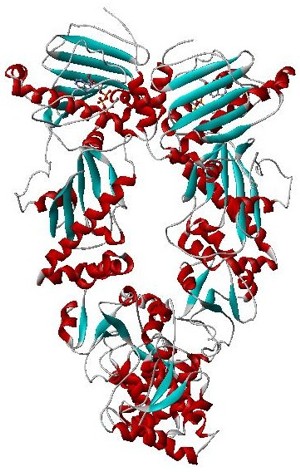
Hsp90 is a chaperone protein that assists other proteins to fold properly, stabilizes proteins against heat stress, and aids in protein degradation. It also stabilizes a number of proteins required for tumor growth, which is why Hsp90 inhibitors are investigated as anti-cancer drugs.
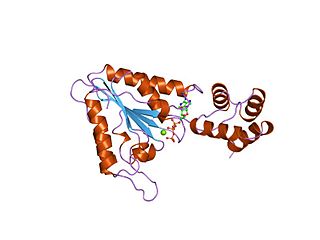
AAA proteins or ATPases Associated with diverse cellular Activities are a protein family sharing a common conserved module of approximately 230 amino acid residues. This is a large, functionally diverse protein family belonging to the AAA+ protein superfamily of ring-shaped P-loop NTPases, which exert their activity through the energy-dependent remodeling or translocation of macromolecules.

Endopeptidase Clp (EC 3.4.21.92, endopeptidase Ti, caseinolytic protease, protease Ti, ATP-dependent Clp protease, ClpP, Clp protease). This enzyme catalyses the following chemical reaction
Non-chaperonin molecular chaperone ATPase (EC 3.6.4.10, molecular chaperone Hsc70 ATPase) is an enzyme with systematic name ATP phosphohydrolase (polypeptide-polymerizing). This enzyme catalyses the following chemical reaction

26S protease regulatory subunit 6A, also known as 26S proteasome AAA-ATPase subunit Rpt5, is an enzyme that in humans is encoded by the PSMC3 gene. This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.
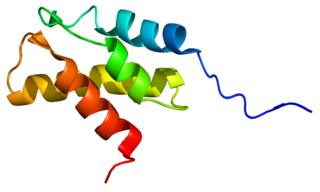
26S protease regulatory subunit 8, also known as 26S proteasome AAA-ATPase subunit Rpt6, is an enzyme that in humans is encoded by the PSMC5 gene. This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.

26S proteasome non-ATPase regulatory subunit 4, also as known as 26S Proteasome Regulatory Subunit Rpn10, is an enzyme that in humans is encoded by the PSMD4 gene. This protein is one of the 19 essential subunits that contributes to the complete assembly of 19S proteasome complex.

26S protease regulatory subunit 7, also known as 26S proteasome AAA-ATPase subunit Rpt1, is an enzyme that in humans is encoded by the PSMC2 gene This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex. Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.

26S protease regulatory subunit 4, also known as 26S proteasome AAA-ATPase subunit Rpt2, is an enzyme that in humans is encoded by the PSMC1 gene. This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex. Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.
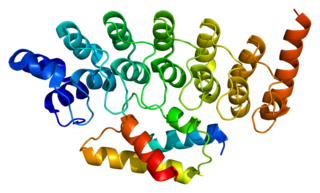
26S protease regulatory subunit 6B, also known as 26S proteasome AAA-ATPase subunit Rpt3, is an enzyme that in humans is encoded by the PSMC4 gene. This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.

Proteasome activator complex subunit 2 is a protein that in humans is encoded by the PSME2 gene.

26S protease regulatory subunit S10B, also known as 26S proteasome AAA-ATPase subunit Rpt4, is an enzyme that in humans is encoded by the PSMC6 gene. This protein is one of the 19 essential subunits of a complete assembled 19S proteasome complex Six 26S proteasome AAA-ATPase subunits together with four non-ATPase subunits form the base sub complex of 19S regulatory particle for proteasome complex.

26S proteasome non-ATPase regulatory subunit 2, also as known as 26S Proteasome Regulatory Subunit Rpn1, is an enzyme that in humans is encoded by the PSMD2 gene.

ATP-dependent Clp protease proteolytic subunit (ClpP) is an enzyme that in humans is encoded by the CLPP gene. This protein is an essential component to form the protein complex of Clp protease.
The N-end rule is a rule that governs the rate of protein degradation through recognition of the N-terminal residue of proteins. The rule states that the N-terminal amino acid of a protein determines its half-life. The rule applies to both eukaryotic and prokaryotic organisms, but with different strength, rules, and outcome. In eukaryotic cells, these N-terminal residues are recognized and targeted by ubiquitin ligases, mediating ubiquitination thereby marking the protein for degradation. The rule was initially discovered by Alexander Varshavsky and co-workers in 1986. However, only rough estimations of protein half-life can be deduced from this 'rule', as N-terminal amino acid modification can lead to variability and anomalies, whilst amino acid impact can also change from organism to organism. Other degradation signals, known as degrons, can also be found in sequence.

ClpS is an N-recognin in the N-end rule pathway. ClpS interacts with protein substrates that have a bulky hydrophobic residue at the N-terminus. The protein substrate is then degraded by the ClpAP protease.
ATP-dependent Clp protease ATP-binding subunit clpX-like, mitochondrial is an enzyme that in humans is encoded by the CLPX gene. This protein is a member of the family of AAA Proteins and is to form the protein complex of Clp protease.
Alfred Lewis Goldberg was an American cell biologist-biochemist and professor at Harvard University. His major discoveries have concerned the mechanisms and physiological importance of protein degradation in cells. Of wide impact have been his lab's demonstration that all cells contain a pathway for selectively eliminating misfolded proteins, his discoveries about the role of proteasomes in this process and of the enzyme systems catalyzing protein breakdown in bacteria, his elucidating the mechanisms for muscle atrophy and the role of proteasomes in antigen presentation to the immune system, and his introduction of proteasome inhibitors now widely used as research tools and in the treatment of blood cancers.















