Related Research Articles

In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA. The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences, and often a substantial fraction of junk DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome.

A transposable element is a nucleic acid sequence in DNA that can change its position within a genome, sometimes creating or reversing mutations and altering the cell's genetic identity and genome size. Transposition often results in duplication of the same genetic material. In the human genome, L1 and Alu elements are two examples. Barbara McClintock's discovery of them earned her a Nobel Prize in 1983. Its importance in personalized medicine is becoming increasingly relevant, as well as gaining more attention in data analytics given the difficulty of analysis in very high dimensional spaces.
Viroids are small single-stranded, circular RNAs that are infectious pathogens. Unlike viruses, they have no protein coating. All known viroids are inhabitants of angiosperms, and most cause diseases, whose respective economic importance to humans varies widely. A recent metatranscriptomics study suggests that the host diversity of viroids and other viroid-like elements is broader than previously thought and that it would not be limited to plants, encompassing even the prokaryotes.
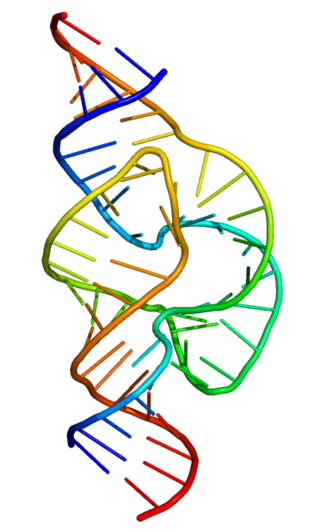
Ribozymes are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demonstrated that RNA can be both genetic material and a biological catalyst, and contributed to the RNA world hypothesis, which suggests that RNA may have been important in the evolution of prebiotic self-replicating systems.
Virusoids are circular single-stranded RNA(s) dependent on viruses for replication and encapsidation. The genome of virusoids consist of several hundred (200–400) nucleotides and does not code for any proteins.

Retrotransposons are a type of genetic component that copy and paste themselves into different genomic locations (transposon) by converting RNA back into DNA through the reverse transcription process using an RNA transposition intermediate.
The Pospiviroidae are a incertae sedis family of ssRNA viroids with 5 genera and 39 species, including the first viroid to be discovered, PSTVd, which is part of genus Pospiviroid. Their secondary structure is key to their biological activity. The classification of this family is based on differences in the conserved central region sequence. Pospiviroidae replication occurs in an asymmetric fashion via host cell RNA polymerase, RNase, and RNA ligase. its hosts are plants, specifically dicotyledons and some monocotyledons
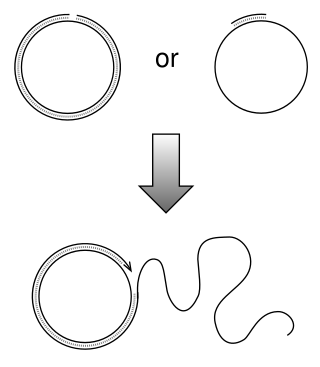

Rolling circle replication (RCR) is a process of unidirectional nucleic acid replication that can rapidly synthesize multiple copies of circular molecules of DNA or RNA, such as plasmids, the genomes of bacteriophages, and the circular RNA genome of viroids. Some eukaryotic viruses also replicate their DNA or RNA via the rolling circle mechanism.
Non-cellular life, also known as acellular life, is life that exists without a cellular structure for at least part of its life cycle. Historically, most definitions of life postulated that an organism must be composed of one or more cells, but this is for some no longer considered necessary, and modern criteria allow for forms of life based on other structural arrangements.
Exon shuffling is a molecular mechanism for the formation of new genes. It is a process through which two or more exons from different genes can be brought together ectopically, or the same exon can be duplicated, to create a new exon-intron structure. There are different mechanisms through which exon shuffling occurs: transposon mediated exon shuffling, crossover during sexual recombination of parental genomes and illegitimate recombination.

Mobile genetic elements (MGEs) sometimes called selfish genetic elements are a type of genetic material that can move around within a genome, or that can be transferred from one species or replicon to another. MGEs are found in all organisms. In humans, approximately 50% of the genome is thought to be MGEs. MGEs play a distinct role in evolution. Gene duplication events can also happen through the mechanism of MGEs. MGEs can also cause mutations in protein coding regions, which alters the protein functions. These mechanisms can also rearrange genes in the host genome generating variation. These mechanism can increase fitness by gaining new or additional functions. An example of MGEs in evolutionary context are that virulence factors and antibiotic resistance genes of MGEs can be transported to share genetic code with neighboring bacteria. However, MGEs can also decrease fitness by introducing disease-causing alleles or mutations. The set of MGEs in an organism is called a mobilome, which is composed of a large number of plasmids, transposons and viruses.

The Avsunviroidae are a family of viroids. There are four species in three genera. They consist of RNA genomes between 246 and 375 nucleotides in length. They are single-stranded covalent circles and have intramolecular base pairing. All members lack a central conserved region.

The hammerhead ribozyme is an RNA motif that catalyzes reversible cleavage and ligation reactions at a specific site within an RNA molecule. It is one of several catalytic RNAs (ribozymes) known to occur in nature. It serves as a model system for research on the structure and properties of RNA, and is used for targeted RNA cleavage experiments, some with proposed therapeutic applications. Named for the resemblance of early secondary structure diagrams to a hammerhead shark, hammerhead ribozymes were originally discovered in two classes of plant virus-like RNAs: satellite RNAs and viroids. They are also known in some classes of retrotransposons, including the retrozymes. The hammerhead ribozyme motif has been ubiquitously reported in lineages across the tree of life.

The hairpin ribozyme is a small section of RNA that can act as a ribozyme. Like the hammerhead ribozyme it is found in RNA satellites of plant viruses. It was first identified in the minus strand of the tobacco ringspot virus (TRSV) satellite RNA where it catalyzes self-cleavage and joining (ligation) reactions to process the products of rolling circle virus replication into linear and circular satellite RNA molecules. The hairpin ribozyme is similar to the hammerhead ribozyme in that it does not require a metal ion for the reaction.

The hepatitis delta virus (HDV) ribozyme is a non-coding RNA found in the hepatitis delta virus that is necessary for viral replication and is the only known human virus that utilizes ribozyme activity to infect its host. The ribozyme acts to process the RNA transcripts to unit lengths in a self-cleavage reaction during replication of the hepatitis delta virus, which is thought to propagate by a double rolling circle mechanism. The ribozyme is active in vivo in the absence of any protein factors and was the fastest known naturally occurring self-cleaving RNA at the time of its discovery.
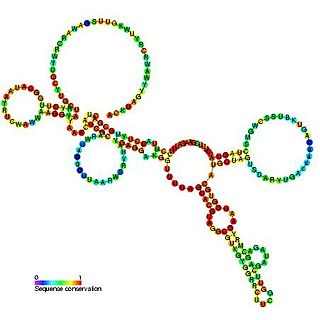
The R2 RNA element is a non-long terminal repeat (non-LTR) retrotransposable element that inserts at a specific site in the 28S ribosomal RNA (rRNA) genes of most insect genomes. In order to insert itself into the genome, retrotransposon encoded protein (R2) protein makes a specific nick in one of the DNA strands at the insertion site and uses the 3′ hydroxyl group exposed by this nick to prime the reverse transcription process termed target primed reverse transcription (TPRT), where the RNA genome is transcribed into DNA.

Circular RNA is a type of single-stranded RNA which, unlike linear RNA, forms a covalently closed continuous loop. In circular RNA, the 3' and 5' ends normally present in an RNA molecule have been joined together. This feature confers numerous properties to circular RNA, many of which have only recently been identified.

Long interspersed nuclear elements (LINEs) are a group of non-LTR retrotransposons that are widespread in the genome of many eukaryotes. LINEs contain an internal Pol II promoter to initiate transcription into mRNA, and encode one or two proteins, ORF1 and ORF2. The functional domains present within ORF1 vary greatly among LINEs, but often exhibit RNA/DNA binding activity. ORF2 is essential to successful retrotransposition, and encodes a protein with both reverse transcriptase and endonuclease activity.

Short interspersed nuclear elements (SINEs) are non-autonomous, non-coding transposable elements (TEs) that are about 100 to 700 base pairs in length. They are a class of retrotransposons, DNA elements that amplify themselves throughout eukaryotic genomes, often through RNA intermediates. SINEs compose about 13% of the mammalian genome.

Ribozyviria is a realm of satellite nucleic acids — infectious agents that resemble viruses, but cannot replicate without a helper virus. Established in ICTV TaxoProp 2020.012D, the realm is named after the presence of genomic and antigenomic ribozymes of the Deltavirus type. The agents in Ribozyviria are satellite nucleic acids, which are distinct from satellite viruses in that they do not encode all of their own structural proteins but require proteins from their helper viruses in order to assemble. Additional common features include a rod-like structure, an RNA-binding "delta antigen" encoded in the genome, and animal hosts. Furthermore, the size range of the genomes of these viruses is between around 1547–1735nt, they encode a hammerhead ribozyme or a hepatitis delta virus ribozyme, and their coding capacity only involves one conserved protein. Most lineages of this realm are poorly understood, the notable exception being the genus Deltavirus, comprising the causal agents of hepatitis D in humans.
References
- 1 2 Cervera, Amelia; Urbina, Denisse; de la Peña, Marcos (2016-06-23). "Retrozymes are a unique family of non-autonomous retrotransposons with hammerhead ribozymes that propagate in plants through circular RNAs". Genome Biology. 17 (1): 135. doi: 10.1186/s13059-016-1002-4 . ISSN 1474-760X. PMC 4918200 . PMID 27339130.
- 1 2 3 4 Cervera, Amelia; Peña, Marcos (2020-05-21). "Small circRNAs with self-cleaving ribozymes are highly expressed in diverse metazoan transcriptomes". Nucleic Acids Research. 48 (9): 5054–5064. doi:10.1093/nar/gkaa187. ISSN 0305-1048. PMC 7229834 . PMID 32198887.
- 1 2 3 4 Lee, Benjamin D.; Koonin, Eugene V. (2022-01-12). "Viroids and Viroid-like Circular RNAs: Do They Descend from Primordial Replicators?". Life. 12 (1): 103. doi: 10.3390/life12010103 . ISSN 2075-1729. PMC 8781251 . PMID 35054497.
- 1 2 de la Peña, Marcos; Cervera, Amelia (2017-08-03). "Circular RNAs with hammerhead ribozymes encoded in eukaryotic genomes: The enemy at home". RNA Biology. 14 (8): 985–991. doi:10.1080/15476286.2017.1321730. ISSN 1547-6286. PMC 5680766 . PMID 28448743.
- ↑ "ViroidDB". viroids.org. Retrieved 2022-01-20.
- 1 2 Lee, Benjamin D; Neri, Uri; Oh, Caleb J; Simmonds, Peter; Koonin, Eugene V (2022-01-07). "ViroidDB: a database of viroids and viroid-like circular RNAs". Nucleic Acids Research. 50 (D1): D432–D438. doi:10.1093/nar/gkab974. ISSN 0305-1048. PMC 8728161 . PMID 34751403.
- ↑ Cervera, Amelia; de la Peña, Marcos (2021), Scarborough, Robert J; Gatignol, Anne (eds.), "Cloning and Detection of Genomic Retrozymes and Their circRNA Intermediates", Ribozymes, Methods in Molecular Biology, New York, NY: Springer US, vol. 2167, pp. 27–44, doi:10.1007/978-1-0716-0716-9_3, ISBN 978-1-0716-0715-2, PMID 32712913, S2CID 220797209 , retrieved 2022-01-20
- ↑ de la Peña, Marcos (2018), Xiao, Junjie (ed.), "Circular RNAs Biogenesis in Eukaryotes Through Self-Cleaving Hammerhead Ribozymes" , Circular RNAs: Biogenesis and Functions, Advances in Experimental Medicine and Biology, Singapore: Springer, vol. 1087, pp. 53–63, doi:10.1007/978-981-13-1426-1_5, ISBN 978-981-13-1426-1, PMID 30259357 , retrieved 2022-01-20
- ↑ Hepojoki, Jussi; Hetzel, Udo; Paraskevopoulou, Sofia; Drosten, Christian; Balazs Harrach; Zerbini, Francisco Murilo; Koonin, Eugene V; Krupovic, Mart; Dolja, Valerian V.; Kuhn, Jens H. (2021). "Create one new realm (Ribozyviria) including one new family (Kolmioviridae) including genus Deltavirus and seven new genera for a total of 15 species". doi:10.13140/RG.2.2.31235.43041.
{{cite journal}}: Cite journal requires|journal=(help) - ↑ Diener, T. O. (1989-12-01). "Circular RNAs: relics of precellular evolution?". Proceedings of the National Academy of Sciences. 86 (23): 9370–9374. Bibcode:1989PNAS...86.9370D. doi: 10.1073/pnas.86.23.9370 . ISSN 0027-8424. PMC 298497 . PMID 2480600.
- 1 2 Diener, Theodor O. (2016). "Viroids: "living fossils" of primordial RNAs?". Biology Direct. 11 (1): 15. doi: 10.1186/s13062-016-0116-7 . ISSN 1745-6150. PMC 4807594 . PMID 27016066.
- ↑ Moelling, Karin; Broecker, Felix (2021-03-28). "Viroids and the Origin of Life". International Journal of Molecular Sciences. 22 (7): 3476. doi: 10.3390/ijms22073476 . ISSN 1422-0067. PMC 8036462 . PMID 33800543.
- ↑ de la Peña, Marcos; Cervera, Amelia (2017-08-03). "Circular RNAs with hammerhead ribozymes encoded in eukaryotic genomes: The enemy at home". RNA Biology. 14 (8): 985–991. doi:10.1080/15476286.2017.1321730. ISSN 1547-6286. PMC 5680766 . PMID 28448743.