| ATP phosphoribosyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
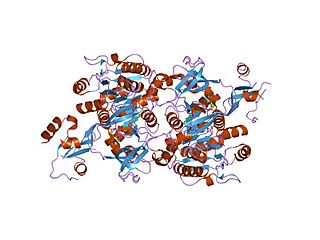
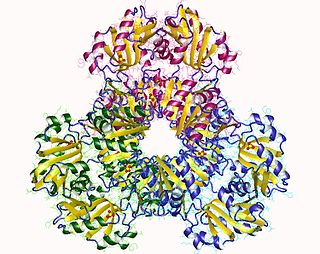
 ATP phosphoribosyltransferase hexamer, Campylobacter jejuni | |||||||||
| Identifiers | |||||||||
| EC no. | 2.4.2.17 | ||||||||
| CAS no. | 9031-46-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| ATP phosphoribosyltransferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 structure of the e.coli atp-phosphoribosyltransferase | |||||||||
| Identifiers | |||||||||
| Symbol | HisG | ||||||||
| Pfam | PF01634 | ||||||||
| Pfam clan | CL0177 | ||||||||
| InterPro | IPR013820 | ||||||||
| PROSITE | PDOC01020 | ||||||||
| SCOP2 | 1nh8 / SCOPe / SUPFAM | ||||||||
| |||||||||
| HisG, C-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
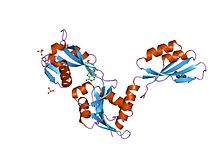 atp phosphoribosyltransferase (atp-prtase) from mycobacterium tuberculosis in complex with amp and histidine | |||||||||
| Identifiers | |||||||||
| Symbol | HisG_C | ||||||||
| Pfam | PF08029 | ||||||||
| Pfam clan | CL0089 | ||||||||
| InterPro | IPR013115 | ||||||||
| |||||||||
In enzymology, an ATP phosphoribosyltransferase (EC 2.4.2.17) is an enzyme that catalyzes the chemical reaction
Contents
- 1-(5-phospho-D-ribosyl)-ATP + diphosphate ATP + 5-phospho-alpha-D-ribose 1-diphosphate
Thus, the two substrates of this enzyme are 1-(5-phospho-D-ribosyl)-ATP and diphosphate, whereas its two products are ATP and 5-phospho-alpha-D-ribose 1-diphosphate.
This enzyme belongs to the family of glycosyltransferases, specifically the pentosyltransferases. The systematic name of this enzyme class is 1-(5-phospho-D-ribosyl)-ATP:diphosphate phospho-alpha-D-ribosyl-transferase. Other names in common use include phosphoribosyl-ATP pyrophosphorylase, adenosine triphosphate phosphoribosyltransferase, phosphoribosyladenosine triphosphate:pyrophosphate, phosphoribosyltransferase, phosphoribosyl ATP synthetase, phosphoribosyl ATP:pyrophosphate phosphoribosyltransferase, phosphoribosyl-ATP:pyrophosphate-phosphoribosyl phosphotransferase, phosphoribosyladenosine triphosphate pyrophosphorylase, and phosphoribosyladenosine triphosphate synthetase.
This enzyme catalyses the first step in the biosynthesis of histidine in bacteria, fungi and plants. It is a member of the larger phosphoribosyltransferase superfamily of enzymes which catalyse the condensation of 5-phospho-alpha-D-ribose 1-diphosphate with nitrogenous bases in the presence of divalent metal ions. [1]
Histidine biosynthesis is an energetically expensive process and ATP phosphoribosyltransferase activity is subject to control at several levels. Transcriptional regulation is based primarily on nutrient conditions and determines the amount of enzyme present in the cell, while feedback inhibition rapidly modulates activity in response to cellular conditions. The enzyme has been shown to be inhibited by 1-(5-phospho-D-ribosyl)-ATP, histidine, ppGpp (a signal associated with adverse environmental conditions) and ADP and AMP (which reflect the overall energy status of the cell). As this pathway of histidine biosynthesis is present only in prokaryotes, plants and fungi, this enzyme is a promising target for the development of novel antimicrobial compounds and herbicides.
ATP phosphoribosyltransferase is found in two distinct forms: a long form containing two catalytic domains and a C-terminal regulatory domain, and a short form in which the regulatory domain is missing. The long form is catalytically competent, but in organisms with the short form, a histidyl-tRNA synthetase paralogue, HisZ, is required for enzyme activity. [2]
The structures of the long form enzymes from Escherichia coli and Mycobacterium tuberculosis have been determined. [3] [4] Interconversion between the various forms is largely reversible and is influenced by the binding of the natural substrates and inhibitors of the enzyme. The two catalytic domains are linked by a two-stranded beta-sheet and together form a "periplasmic binding protein fold". A crevice between these domains contains the active site. The C-terminal domain is not directly involved in catalysis but appears to be involved the formation of hexamers, induced by the binding of inhibitors such as histidine to the enzyme, thus regulating activity.