
Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein folds into its three dimensional tertiary structure.

The Rossmann fold is a tertiary fold found in proteins that bind nucleotides, such as enzyme cofactors FAD, NAD+, and NADP+. This fold is composed of alternating beta strands and alpha helical segments where the beta strands are hydrogen bonded to each other forming an extended beta sheet and the alpha helices surround both faces of the sheet to produce a three-layered sandwich. The classical Rossmann fold contains six beta strands whereas Rossmann-like folds, sometimes referred to as Rossmannoid folds, contain only five strands. The initial beta-alpha-beta (bab) fold is the most conserved segment of the Rossmann fold. The motif is named after Michael Rossmann who first noticed this structural motif in the enzyme lactate dehydrogenase in 1970 and who later observed that this was a frequently occurring motif in nucleotide binding proteins.

Protein tyrosine phosphatases (EC 3.1.3.48, systematic name protein-tyrosine-phosphate phosphohydrolase) are a group of enzymes that remove phosphate groups from phosphorylated tyrosine residues on proteins:

The TIM barrel, also known as an alpha/beta barrel, is a conserved protein fold consisting of eight alpha helices (α-helices) and eight parallel beta strands (β-strands) that alternate along the peptide backbone. The structure is named after triose-phosphate isomerase, a conserved metabolic enzyme. TIM barrels are ubiquitous, with approximately 10% of all enzymes adopting this fold. Further, five of seven enzyme commission (EC) enzyme classes include TIM barrel proteins. The TIM barrel fold is evolutionarily ancient, with many of its members possessing little similarity today, instead falling within the twilight zone of sequence similarity.

Transaldolase is an enzyme of the non-oxidative phase of the pentose phosphate pathway. In humans, transaldolase is encoded by the TALDO1 gene.

Guanosine monophosphate synthetase, also known as GMPS is an enzyme that converts xanthosine monophosphate to guanosine monophosphate.

Cystathionine beta-lyase, also commonly referred to as CBL or β-cystathionase, is an enzyme that primarily catalyzes the following α,β-elimination reaction
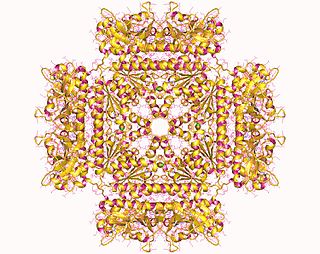

In molecular biology, the protein domain SAICAR synthase is an enzyme which catalyses a reaction to create SAICAR. In enzymology, this enzyme is also known as phosphoribosylaminoimidazolesuccinocarboxamide synthase. It is an enzyme that catalyzes the chemical reaction

In Enzymology, a dUTP diphosphatase (EC 3.6.1.23) is an enzyme that catalyzes the chemical reaction
Exopolyphosphatase (PPX) is a phosphatase enzyme which catalyzes the hydrolysis of inorganic polyphosphate, a linear molecule composed of up to 1000 or more monomers linked by phospho-anhydride bonds. PPX is a processive exophosphatase, which means that it begins at the ends of the polyphosphate chain and cleaves the phospho-anhydride bonds to release orthophosphate as it moves along the polyphosphate molecule. PPX has several characteristics which distinguish it from other known polyphosphatases, namely that it does not act on ATP, has a strong preference for long chain polyphosphate, and has a very low affinity for polyphosphate molecules with less than 15 phosphate monomers.
In enzymology, an undecaprenyl-diphosphatase (EC 3.6.1.27) is an enzyme that catalyzes the chemical reaction
In enzymology, an IMP cyclohydrolase (EC 3.5.4.10) is an enzyme that catalyzes the chemical reaction

The alpha/beta hydrolase superfamily is a superfamily of hydrolytic enzymes of widely differing phylogenetic origin and catalytic function that share a common fold. The core of each enzyme is an alpha/beta-sheet, containing 8 beta strands connected by 6 alpha helices. The enzymes are believed to have diverged from a common ancestor, retaining little obvious sequence similarity, but preserving the arrangement of the catalytic residues. All have a catalytic triad, the elements of which are borne on loops, which are the best-conserved structural features of the fold.
Aminoglycoside-3'-phosphotransferase, also known as aminoglycoside kinase, is an enzyme that primarily catalyzes the addition of phosphate from ATP to the 3'-hydroxyl group of a 4,6-disubstituted aminoglycoside, such as kanamycin. However, APH(3') has also been found to phosphorylate at the 5'-hydroxyl group in 4,5-disubstituted aminoglycosides, which lack a 3'-hydroxyl group, and to diphosphorylate hydroxyl groups in aminoglycosides that have both 3'- and 5'-hydroxyl groups. Primarily positively charged at biological conditions, aminoglycosides bind to the negatively charged backbone of nucleic acids to disrupt protein synthesis, effectively inhibiting bacterial cell growth. APH(3') mediated phosphorylation of aminoglycosides effectively disrupts their mechanism of action, introducing a phosphate group that reduces their binding affinity due to steric hindrances and unfavorable electrostatic interactions. APH(3') is primarily found in certain species of gram-positive bacteria.

Acylphosphatase-2 is an enzyme that in humans is encoded by the ACYP2 gene.

Acylphosphatase-1 is an enzyme that in humans is encoded by the ACYP1 gene.
The Walker A and Walker B motifs are protein sequence motifs, known to have highly conserved three-dimensional structures. These were first reported in ATP-binding proteins by Walker and co-workers in 1982.

In molecular biology, the protein domain surE refers to survival protein E. It was originally found that cells that did not contain this protein, could not survive in the stationary phase, at above normal temperatures, and in high-salt media. Hence the name, survival protein E. It is a metal ion-dependent phosphatase that is found in bacteria, and eukaryotes. It is an important stress response protein. This domain is found in acid phosphatases, 5'-nucleotidases, 3'-nucleotidases and exopolyphosphatases.

In molecular biology, YopH, N-terminal refers to an evolutionary conserved protein domain. This entry represents the N-terminal domain of YopH protein tyrosine phosphatase (PTP).


















