A protein phosphatase is a phosphatase enzyme that removes a phosphate group from the phosphorylated amino acid residue of its substrate protein. Protein phosphorylation is one of the most common forms of reversible protein posttranslational modification (PTM), with up to 30% of all proteins being phosphorylated at any given time. Protein kinases (PKs) are the effectors of phosphorylation and catalyse the transfer of a γ-phosphate from ATP to specific amino acids on proteins. Several hundred PKs exist in mammals and are classified into distinct super-families. Proteins are phosphorylated predominantly on Ser, Thr and Tyr residues, which account for 79.3, 16.9 and 3.8% respectively of the phosphoproteome, at least in mammals. In contrast, protein phosphatases (PPs) are the primary effectors of dephosphorylation and can be grouped into three main classes based on sequence, structure and catalytic function. The largest class of PPs is the phosphoprotein phosphatase (PPP) family comprising PP1, PP2A, PP2B, PP4, PP5, PP6 and PP7, and the protein phosphatase Mg2+- or Mn2+-dependent (PPM) family, composed primarily of PP2C. The protein Tyr phosphatase (PTP) super-family forms the second group, and the aspartate-based protein phosphatases the third. The protein pseudophosphatases form part of the larger phosphatase family, and in most cases are thought to be catalytically inert, instead functioning as phosphate-binding proteins, integrators of signalling or subcellular traps. Examples of membrane-spanning protein phosphatases containing both active (phosphatase) and inactive (pseudophosphatase) domains linked in tandem are known, conceptually similar to the kinase and pseudokinase domain polypeptide structure of the JAK pseudokinases. A complete comparative analysis of human phosphatases and pseudophosphatases has been completed by Manning and colleagues, forming a companion piece to the ground-breaking analysis of the human kinome, which encodes the complete set of ~536 human protein kinases.

In biochemistry, a phosphatase is an enzyme that uses water to cleave a phosphoric acid monoester into a phosphate ion and an alcohol. Because a phosphatase enzyme catalyzes the hydrolysis of its substrate, it is a subcategory of hydrolases. Phosphatase enzymes are essential to many biological functions, because phosphorylation and dephosphorylation serve diverse roles in cellular regulation and signaling. Whereas phosphatases remove phosphate groups from molecules, kinases catalyze the transfer of phosphate groups to molecules from ATP. Together, kinases and phosphatases direct a form of post-translational modification that is essential to the cell's regulatory network.

The enzyme alkaline phosphatase has the physiological role of dephosphorylating compounds. The enzyme is found across a multitude of organisms, prokaryotes and eukaryotes alike, with the same general function but in different structural forms suitable to the environment they function in. Alkaline phosphatase is found in the periplasmic space of E. coli bacteria. This enzyme is heat stable and has its maximum activity at high pH. In humans, it is found in many forms depending on its origin within the body – it plays an integral role in metabolism within the liver and development within the skeleton. Due to its widespread prevalence in these areas, its concentration in the bloodstream is used by diagnosticians as a biomarker in helping determine diagnoses such as hepatitis or osteomalacia.
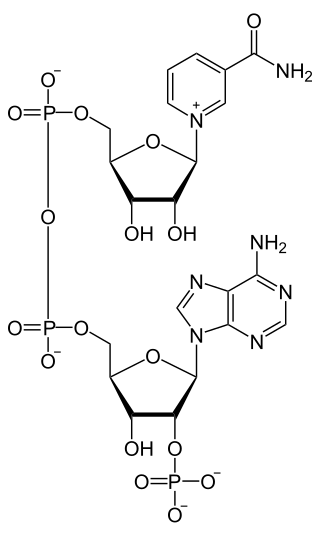
Nicotinamide adenine dinucleotide phosphate, abbreviated NADP+ or, in older notation, TPN (triphosphopyridine nucleotide), is a cofactor used in anabolic reactions, such as the Calvin cycle and lipid and nucleic acid syntheses, which require NADPH as a reducing agent ('hydrogen source'). NADPH is the reduced form of NADP+, the oxidized form. NADP+ is used by all forms of cellular life.

In biochemistry, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate+hydrogen) to an acceptor.
Phosphofructokinase-2 (6-phosphofructo-2-kinase, PFK-2) or fructose bisphosphatase-2 (FBPase-2), is an enzyme indirectly responsible for regulating the rates of glycolysis and gluconeogenesis in cells. It catalyzes formation and degradation of a significant allosteric regulator, fructose-2,6-bisphosphate (Fru-2,6-P2) from substrate fructose-6-phosphate. Fru-2,6-P2 contributes to the rate-determining step of glycolysis as it activates enzyme phosphofructokinase 1 in the glycolysis pathway, and inhibits fructose-1,6-bisphosphatase 1 in gluconeogenesis. Since Fru-2,6-P2 differentially regulates glycolysis and gluconeogenesis, it can act as a key signal to switch between the opposing pathways. Because PFK-2 produces Fru-2,6-P2 in response to hormonal signaling, metabolism can be more sensitively and efficiently controlled to align with the organism's glycolytic needs. This enzyme participates in fructose and mannose metabolism. The enzyme is important in the regulation of hepatic carbohydrate metabolism and is found in greatest quantities in the liver, kidney and heart. In mammals, several genes often encode different isoforms, each of which differs in its tissue distribution and enzymatic activity. The family described here bears a resemblance to the ATP-driven phospho-fructokinases, however, they share little sequence similarity, although a few residues seem key to their interaction with fructose 6-phosphate.

The enzyme glucose 6-phosphatase (EC 3.1.3.9, G6Pase; systematic name D-glucose-6-phosphate phosphohydrolase) catalyzes the hydrolysis of glucose 6-phosphate, resulting in the creation of a phosphate group and free glucose:

Protein tyrosine phosphatases (EC 3.1.3.48, systematic name protein-tyrosine-phosphate phosphohydrolase) are a group of enzymes that remove phosphate groups from phosphorylated tyrosine residues on proteins:
An esterase is a hydrolase enzyme that splits esters into an acid and an alcohol in a chemical reaction with water called hydrolysis.
Acid phosphatase is an enzyme that frees attached phosphoryl groups from other molecules during digestion. It can be further classified as a phosphomonoesterase. It is stored in lysosomes and functions when these fuse with endosomes, which are acidified while they function; therefore, it has an acid pH optimum. This enzyme is present in many animal and plant species.

Phosphoglycerate mutase (PGM) is any enzyme that catalyzes step 8 of glycolysis - the internal transfer of a phosphate group from C-3 to C-2 which results in the conversion of 3-phosphoglycerate (3PG) to 2-phosphoglycerate (2PG) through a 2,3-bisphosphoglycerate intermediate. These enzymes are categorized into the two distinct classes of either cofactor-dependent (dPGM) or cofactor-independent (iPGM). The dPGM enzyme is composed of approximately 250 amino acids and is found in all vertebrates as well as in some invertebrates, fungi, and bacteria. The iPGM class is found in all plants and algae as well as in some invertebrate, fungi, and Gram-positive bacteria. This class of PGM enzyme shares the same superfamily as alkaline phosphatase.

In enzymology, aldose reductase is a cytosolic NADPH-dependent oxidoreductase that catalyzes the reduction of a variety of aldehydes and carbonyls, including monosaccharides. It is primarily known for catalyzing the reduction of glucose to sorbitol, the first step in polyol pathway of glucose metabolism.
In enzymology, a sorbitol-6-phosphate dehydrogenase (EC 1.1.1.140) is an enzyme that catalyzes the chemical reaction
In enzymology, an aldose-6-phosphate reductase (NADPH) (EC 1.1.1.200) is an enzyme that catalyzes the chemical reaction

In enzymology, a nucleoside-diphosphatase (EC 3.6.1.6) is an enzyme that catalyzes the chemical reaction

The enzyme Inositol phosphate-phosphatase is of the phosphodiesterase family of enzymes. It is involved in the phosphophatidylinositol signaling pathway, which affects a wide array of cell functions, including but not limited to, cell growth, apoptosis, secretion, and information processing. Inhibition of inositol monophosphatase may be key in the action of lithium in treating bipolar disorder, specifically manic depression.
The enzyme pyridoxal phosphatase (EC 3.1.3.74) catalyzes the reaction
The enzyme sucrose-phosphatase (EC 3.1.3.24) catalyzes the reaction
In enzymology, a [isocitrate dehydrogenase (NADP+)] kinase (EC 2.7.11.5) is an enzyme that catalyzes the chemical reaction:
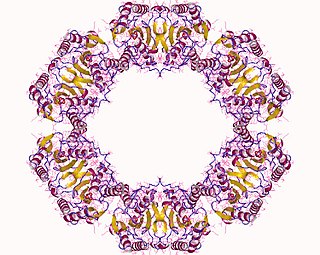

The enzyme protein serine/threonine phosphatase is a form of phosphoprotein phosphatase that acts upon phosphorylated serine/threonine residues:













