
Pseudomonadota is a major phylum of Gram-negative bacteria. The renaming of several prokaryote phyla in 2021, including Pseudomonadota, remains controversial among microbiologists, many of whom continue to use the earlier name Proteobacteria, of long standing in the literature. The phylum Proteobacteria includes a wide variety of pathogenic genera, such as Escherichia, Salmonella, Vibrio, Yersinia, Legionella, and many others. Others are free-living (non-parasitic) and include many of the bacteria responsible for nitrogen fixation.
The Aquificota phylum is a diverse collection of bacteria that live in harsh environmental settings. The name Aquificota was given to this phylum based on an early genus identified within this group, Aquifex, which is able to produce water by oxidizing hydrogen. They have been found in springs, pools, and oceans. They are autotrophs, and are the primary carbon fixers in their environments. These bacteria are Gram-negative, non-spore-forming rods. They are true bacteria as opposed to the other inhabitants of extreme environments, the Archaea.
The Chloroflexia are a class of bacteria in the phylum Chloroflexota. Chloroflexia are typically filamentous, and can move about through bacterial gliding. It is named after the order Chloroflexales.
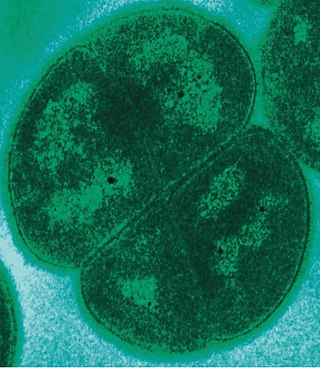
Deinococcota is a phylum of bacteria with a single class, Deinococci, that are highly resistant to environmental hazards, also known as extremophiles. These bacteria have thick cell walls that give them gram-positive stains, but they include a second membrane and so are closer in structure to those of gram-negative bacteria.

The Chlamydiota are a bacterial phylum and class whose members are remarkably diverse, including pathogens of humans and animals, symbionts of ubiquitous protozoa, and marine sediment forms not yet well understood. All of the Chlamydiota that humans have known about for many decades are obligate intracellular bacteria; in 2020 many additional Chlamydiota were discovered in ocean-floor environments, and it is not yet known whether they all have hosts. Historically it was believed that all Chlamydiota had a peptidoglycan-free cell wall, but studies in the 2010s demonstrated a detectable presence of peptidoglycan, as well as other important proteins.

The Pasteurellaceae comprise a large family of Gram-negative bacteria. Most members live as commensals on mucosal surfaces of birds and mammals, especially in the upper respiratory tract. Pasteurellaceae are typically rod-shaped, and are a notable group of facultative anaerobes. Their biochemical characteristics can be distinguished from the related Enterobacteriaceae by the presence of oxidase, and from most other similar bacteria by the absence of flagella.
Xanthomonadaceae is a family of Pseudomonadota within the Xanthomonadales order. It was previously known as Lysobacteraceae.
The Thermotogota are a phylum of the domain Bacteria. The phylum Thermotogota is composed of Gram-negative staining, anaerobic, and mostly thermophilic and hyperthermophilic bacteria.

Streptomycetaceae is a family of Actinomycetota, making up the monotypic order Streptomycetales. It includes the important genus Streptomyces. This was the original source of many antibiotics, namely streptomycin, the first antibiotic against tuberculosis.

Alphaproteobacteria is a class of bacteria in the phylum Pseudomonadota. The Magnetococcales and Mariprofundales are considered basal or sister to the Alphaproteobacteria. The Alphaproteobacteria are highly diverse and possess few commonalities, but nevertheless share a common ancestor. Like all Proteobacteria, its members are gram-negative and some of its intracellular parasitic members lack peptidoglycan and are consequently gram variable.
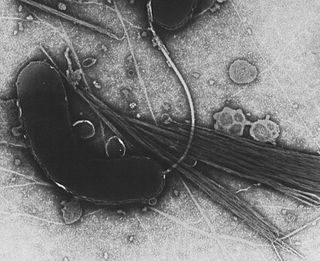
Gammaproteobacteria is a class of bacteria in the phylum Pseudomonadota. It contains about 250 genera, which makes it the most genus-rich taxon of the Prokaryotes. Several medically, ecologically, and scientifically important groups of bacteria belong to this class. It is composed by all Gram-negative microbes and is the most phylogenetically and physiologically diverse class of Proteobacteria.

Stenotrophomonas is a genus of Gram-negative bacteria, comprising at least ten species. The main reservoirs of Stenotrophomonas are soil and plants. Stenotrophomonas species range from common soil organisms to opportunistic human pathogens ; the molecular taxonomy of the genus is still somewhat unclear.

Xanthomonas is a genus of bacteria, many of which cause plant diseases. There are at least 27 plant associated Xanthomonas spp., that all together infect at least 400 plant species. Different species typically have specific host and/or tissue range and colonization strategies.

The genus Lysobacter belongs to the family Xanthomonadaceae within the Gammaproteobacteria and includes at least 46 named species, including: Lysobacter enzymogenes, L. antibioticus, L. gummosus, L. brunescens, L. defluvii, L. niabensis, L. niastensis, L. daejeonensis, L. yangpyeongensis, L. koreensis, L. concretionis, L. spongiicola, and L. capsici. Lysobacter spp. were originally grouped with myxobacteria because they shared the distinctive trait of gliding motility, but they uniquely display a number of traits that distinguish them from other taxonomically and ecologically related microbes including high genomic G+C content and the lack of flagella. The feature of gliding motility alone has piqued the interest of many, since the role of gliding bacteria in soil ecology is poorly understood. In addition, while a number of different mechanisms have been proposed for gliding motility among a wide range of bacterial species, the genetic mechanism in Lysobacter remains unknown. Members of the Lysobacter group have gained broad interest for production of extracellular enzymes. The group is also regarded as a rich source for production of novel antibiotics, such as β-lactams containing substituted side chains, macrocyclic lactams and macrocyclic peptide or depsipeptide antibiotics like the katanosins.
The Synergistota is a phylum of anaerobic bacteria that show Gram-negative staining and have rod/vibrioid cell shape. Although Synergistota have a diderm cell envelope, the genes for various proteins involved in lipopolysaccharides biosynthesis have not yet been detected in Synergistota, indicating that they may have an atypical outer cell envelope. The Synergistota inhabit a majority of anaerobic environments including animal gastrointestinal tracts, soil, oil wells, and wastewater treatment plants and they are also present in sites of human diseases such as cysts, abscesses, and areas of periodontal disease. Due to their presence at illness related sites, the Synergistota are suggested to be opportunistic pathogens but they can also be found in healthy individuals in the microbiome of the umbilicus and in normal vaginal flora. Species within this phylum have also been implicated in periodontal disease, gastrointestinal infections and soft tissue infections. Other species from this phylum have been identified as significant contributors in the degradation of sludge for production of biogas in anaerobic digesters and are potential candidates for use in renewable energy production through their production of hydrogen gas. All of the known Synergistota species and genera are presently part of a single class (Synergistia), order (Synergistiales), and family (Synergistaceae).
The Negativicutes are a class of bacteria in the phylum Bacillota, whose members have a peculiar cell wall with a lipopolysaccharide outer membrane which stains gram-negative, unlike most other members of the Bacillota. Although several neighbouring Clostridia species also stain gram-negative, the proteins responsible for the unusual diderm structure of the Negativicutes may have actually been laterally acquired from Pseudomonadota. Additional research is required to confirm the origin of the diderm cell envelope in the Negativicutes.
Pseudoxanthomonas is a genus of Gram-negative bacteria in the family Xanthomonadaceae from the phylum Pseudomonadota. This genus is closely related phylogenetically with the genera Xanthomonas, Xylella, and Stenotrophomonas. The genus was first distinguished in 2000 in biofilter samples, and was later emended by Lee et al. Some of the species in this genus are: P. mexicana, P. japonensis, P. koreensis, P. daejeonensis, and the type species P. broegbernensis.
The Coriobacteriia are a class of Gram-positive bacteria within the Actinomycetota phylum. Species within this group are nonsporulating, strict or facultative anaerobes that are capable of thriving in a diverse set of ecological niches. Gordonibacter species are the only members capable of motility by means of flagella within the class. Several species within the Coriobacteriia class have been implicated with human diseases that range in severity. Atopobium, Olsenella, and Cryptobacterium species have responsible for human oral infections including periodontitis, halitosis, and other endodontic infections. Eggerthella species have been associated with severe blood bacteraemia and ulcerative colitis.

The Yersiniaceae are a family of Gram-negative bacteria that includes some familiar pathogens. For example, the type genus Yersinia includes Yersinia pestis, the causative agent of plague. This family is a member of the order Enterobacterales in the class Gammaproteobacteria of the phylum Pseudomonadota.
The Hafniaceae are a family of Gram-negative bacteria. This family is a member of the order Enterobacterales in the class Gammaproteobacteria of the phylum Pseudomonadota. Genera in this family include the type genus Hafnia, along with Edwardsiella and Obesumbacterium.











