
Protein biosynthesis is a core biological process, occurring inside cells, balancing the loss of cellular proteins through the production of new proteins. Proteins perform a number of critical functions as enzymes, structural proteins or hormones. Protein synthesis is a very similar process for both prokaryotes and eukaryotes but there are some distinct differences.

Adenosine monophosphate (AMP), also known as 5'-adenylic acid, is a nucleotide. AMP consists of a phosphate group, the sugar ribose, and the nucleobase adenine. It is an ester of phosphoric acid and the nucleoside adenosine. As a substituent it takes the form of the prefix adenylyl-.

Nicotinamide adenine dinucleotide (NAD) is a coenzyme central to metabolism. Found in all living cells, NAD is called a dinucleotide because it consists of two nucleotides joined through their phosphate groups. One nucleotide contains an adenine nucleobase and the other, nicotinamide. NAD exists in two forms: an oxidized and reduced form, abbreviated as NAD+ and NADH (H for hydrogen), respectively.
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In eukaryotes, polyadenylation is part of the process that produces mature mRNA for translation. In many bacteria, the poly(A) tail promotes degradation of the mRNA. It, therefore, forms part of the larger process of gene expression.
A nucleoside triphosphate is a nucleoside containing a nitrogenous base bound to a 5-carbon sugar, with three phosphate groups bound to the sugar. They are the molecular precursors of both DNA and RNA, which are chains of nucleotides made through the processes of DNA replication and transcription. Nucleoside triphosphates also serve as a source of energy for cellular reactions and are involved in signalling pathways.

Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease.
The adenylate energy charge is an index used to measure the energy status of biological cells.

Transcriptional modification or co-transcriptional modification is a set of biological processes common to most eukaryotic cells by which an RNA primary transcript is chemically altered following transcription from a gene to produce a mature, functional RNA molecule that can then leave the nucleus and perform any of a variety of different functions in the cell. There are many types of post-transcriptional modifications achieved through a diverse class of molecular mechanisms.

Polynucleotide Phosphorylase (PNPase) is a bifunctional enzyme with a phosphorolytic 3' to 5' exoribonuclease activity and a 3'-terminal oligonucleotide polymerase activity. That is, it dismantles the RNA chain starting at the 3' end and working toward the 5' end. It also synthesizes long, highly heteropolymeric tails in vivo. It accounts for all of the observed residual polyadenylation in strains of Escherichia coli missing the normal polyadenylation enzyme. Discovered by Marianne Grunberg-Manago working in Severo Ochoa's lab in 1955, the RNA-polymerization activity of PNPase was initially believed to be responsible for DNA-dependent synthesis of messenger RNA, a notion that got disproved by the late 1950s.
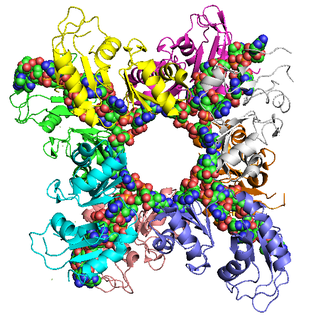
Poly(A)-binding protein is an RNA-binding protein which triggers the binding of eukaryotic initiation factor 4 complex (eIF4G) directly to the poly(A) tail of mRNA which is 200-250 nucleotides long. The poly(A) tail is located on the 3' end of mRNA and was discovered by Mary Edmonds, who also characterized the poly-A polymerase enzyme that generates the poly(a) tail. The binding protein is also involved in mRNA precursors by helping polyadenylate polymerase add the poly(A) nucleotide tail to the pre-mRNA before translation. The nuclear isoform selectively binds to around 50 nucleotides and stimulates the activity of polyadenylate polymerase by increasing its affinity towards RNA. Poly(A)-binding protein is also present during stages of mRNA metabolism including nonsense-mediated decay and nucleocytoplasmic trafficking. The poly(A)-binding protein may also protect the tail from degradation and regulate mRNA production. Without these two proteins in-tandem, then the poly(A) tail would not be added and the RNA would degrade quickly.
Deoxyribonuclease IV (phage-T4-induced) is catalyzes the degradation nucleotides in DsDNA by attacking the 5'-terminal end.
In enzymology, a [glutamate—ammonia-ligase] adenylyltransferase is an enzyme that catalyzes the chemical reaction
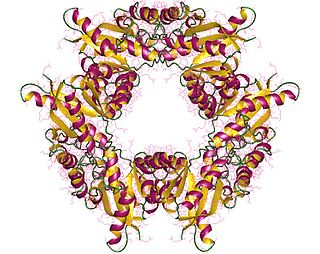

In enzymology, nicotinamide-nucleotide adenylyltransferase (NMNAT) (EC 2.7.7.1) are enzymes that catalyzes the chemical reaction

In enzymology, a sulfate adenylyltransferase is an enzyme that catalyzes the chemical reaction
CCA tRNA nucleotidyltransferase is an enzyme with systematic name CTP,CTP,ATP:tRNA cytidylyl,cytidylyl,adenylyltransferase. This enzyme catalyses the following chemical reaction
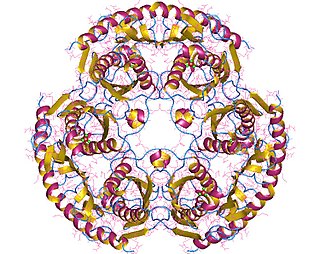
In enzymology, a tRNA nucleotidyltransferase is an enzyme that catalyzes the chemical reaction

Poly(A) polymerase alpha is an enzyme that in humans is encoded by the PAPOLA gene.

The prokaryotic riboflavin biosynthesis protein is a bifunctional enzyme found in bacteria that catalyzes the phosphorylation of riboflavin into flavin mononucleotide (FMN) and the adenylylation of FMN into flavin adenine dinucleotide (FAD). It consists of a C-terminal riboflavin kinase and an N-terminal FMN-adenylyltransferase. This bacterial protein is functionally similar to the monofunctional riboflavin kinases and FMN-adenylyltransferases of eukaryotic organisms, but only the riboflavin kinases are structurally homologous.

GLD-2 is a cytoplasmic poly(A) polymerase (cytoPAPs) which adds successive AMP monomers to the 3’ end of specific RNAs, forming a poly(A) tail, which is a process known as polyadenylation.
Mary P. Edmonds was an American biochemist who made key discoveries regarding the processing of messenger RNA (mRNA). She spent most of her career at the University of Pittsburgh.














