Role of chlorophyllase in chlorophyll breakdown
Of high importance to all photosynthetic organisms is chlorophyll, and so, its synthesis and breakdown are closely regulated throughout the entire life cycle of the plant. Chlorophyll breakdown is most evident in seasonal changes as the plants lose their green color in the autumn; it is also evident in fruit ripening, leaf senescence and flowering. In this first step, chlorophyllase initiates the catabolism of chlorophyll to form chlorophyllide. Chlorophyll degradation occurs in the turnover of chlorophyll, as well as in the event of cell death caused by injuries, pathogenic attack, and other external factors.
Chlorophyllase's role is two-fold as it functions in both de-greening processes, such as autumnal coloration, and is also thought to be involved in turnover and homeostasis of chlorophylls. Chlorophyllase catalysis of the initial step of chlorophyll breakdown is important for plant development and survival. The breakdown serves as a prerequisite in the detoxification of the potentially phototoxic chlorophyll and chlorophyll intermediates as it accompanies leaf senescence to non-fluorescent catabolites. Rapid degradation of chlorophyll and its intermediates is therefore necessary to prevent cell damage due to the potential phototoxicity of chlorophyll. [4] [5] [6]
Reaction and mechanism catalyzed by chlorophyllase
Chlorophyllase catalyzes the hydrolysis of ester bond to yield chlorophyllide and phytol. It reacts via transesterification or hydrolysis of a carboxylic ester in which its natural substrates are 13-OH-chlorophyll a, bacteriochlorophyll and chlorophyll a.
Hydrolysis of chlorophyll starts with the attack of a carbonyl group of chlorophyll by the oxygen of the hydroxyl group of the crucial serine residue of the chlorophyllase. This attack forms a tetrahedral transition state. The double bond of the attacked carbonyl reforms and the serine is then esterified to chlorophyllide. The phytol group consequently leaves the compound and replaces the serine residue on the chlorophyllase enzyme. The addition of water to the reaction cleaves the phytol off the enzyme. Next, through the reverse reaction, the oxygen on the hydroxy group from the water in the previous step attacks the carbonyl of the intermediate in order to form another tetrahedral transition state. The double bond of the carbonyl forms again and the serine residue returns to chlorophyllase and the ester of the chlorophyll is now a carboxylic acid. This product is chlorophyllide. [7]
Chlorophyllide is then broken down to Pheophorbide a. After Pheophorbide a is formed, the porphyrin ring is cleaved by Pheophorbide a oxygenase (PAO) to form RCC (red chlorophyll catabolite) causing the plant to lose its green color. RCC is then broken down into pFCC. [8]
Regulation
Posttranslational Regulation
Citrus sinesis and Chenopodium album were the first plants from which the genes encoding chlorophyllase were isolated. These experiments revealed an uncharacteristic encoded sequence (21 amino acids in Citrus sinensis and 30 amino acids in Chenopodium album) located on the N-terminal that was absent from the mature protein. The chlorophyllase enzyme is a smart choice as the rate limiting enzyme of the catabolic pathway since degreening and the expression of chlorophyllase is induced in ethylene-treated Citrus. Recent data, however, suggests that chlorophyllase is expressed at low levels during natural fruit development, when chlorophyll catabolism usually takes place. Also, some data suggests that chlorophyllase activity is not consistent with degreening during natural senescence. Finally, there is evidence that chlorophyllase has been found in the inner envelope membrane of chloroplast where it does not come in contact with chlorophyll. Recent studies inspired by inconsistent data revealed that chlorophyllase in Citrus lacking the 21 amino sequence on the N-terminal results in extensive chlorophyll breakdown and the degreening effect that should occur in vivo. This cleavage occurs in the chloroplast membrane fraction. Both the full chlorophyllase and the cleaved, mature chlorophyllase, however, experienced similar levels of activity in an in vitro assay. This data suggests that the mature protein comes in contact with its substrate more readily because of the N-terminal sequence and some natural regulation occurs that directly affects enzyme activity. Another possibility is that the suborganelle compartments breaking down allowing a greater amount of enzyme activity. [9]
Chlorophyllide, the product of the reaction catalyzed by chlorophyllase, spontaneously combines with plant lipids such as phosphatidylcholine liposomes along with sulfoquinovosyl diacylglycerol. These two lipids cooperatively inhibit the activity of chlorophyllase, but this inhibition can be reversed by the presence of Mg++, a divalent cation. [10] The activity of chlorophyllase also depends on the pH and ionic content of the medium. The values of kcat and kcat/Km of chlorophyllase in the presence of chlorophyll showed pKa values of 6.3 and 6.7, respectively. Temperature also affects chlorophyllase activity. Wheat chlorophyllase is active from 25 to 75 °C. The enzyme is inactivated at temperatures above 85 °C. Wheat chlorophyllase is stable 20 °C higher than other chlorophyllases. These other chlorophyllases can stay active at temperatures up to 55 °C. [11]
Ethylene induces the synthesis of chlorophyllase and promotes the degreening of citrus fruits. Chlorophyllase was detected in protein extracts of ethylene treated fruit. Ethylene treated fruits had chlorophyllase activity increased by 5 fold in 24 hours. Ethylene, more specifically, induces increased rates of transcription of the chlorophyllase gene. [12] [13]
There is also evidence of a highly conserved serine lipase domain in the chlorophyllase enzyme that contains a serine residue that is essential for enzyme activity. Histidne and aspartic acid residues are also a part of the catalytic triad of chlorophyllase as a serine hydrolase. Specific inhibitors for the serine hydrolase mechanism, therefore, effectively inhibit the chlorophyllase enzyme. Also, mutations at these specific amino acid residues causes complete loss of function since the mutations change the catalytic site of the chlorophyllase enzyme. [7]

Chlorophyll is any of several related green pigments found in cyanobacteria and in the chloroplasts of algae and plants. Its name is derived from the Greek words χλωρός, khloros and φύλλον, phyllon ("leaf"). Chlorophyll allow plants to absorb energy from light.
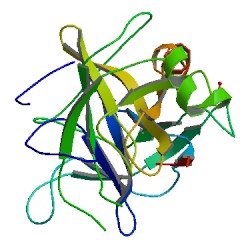
Chymotrypsin (EC 3.4.21.1, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid (tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly those containing leucine at the P1 position.

Proteolysis is the breakdown of proteins into smaller polypeptides or amino acids. Uncatalysed, the hydrolysis of peptide bonds is extremely slow, taking hundreds of years. Proteolysis is typically catalysed by cellular enzymes called proteases, but may also occur by intra-molecular digestion.

A protease is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism, and cell signaling.
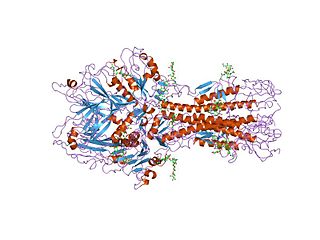
Hemagglutinin esterase (HEs) is a glycoprotein that certain enveloped viruses possess and use as an invading mechanism. HEs helps in the attachment and destruction of certain sialic acid receptors that are found on the host cell surface. Viruses that possess HEs include influenza C virus, toroviruses, and coronaviruses of the subgenus Embecovirus. HEs is a dimer transmembrane protein consisting of two monomers, each monomer is made of three domains. The three domains are: membrane fusion, esterase, and receptor binding domains.

A catalytic triad is a set of three coordinated amino acids that can be found in the active site of some enzymes. Catalytic triads are most commonly found in hydrolase and transferase enzymes. An acid-base-nucleophile triad is a common motif for generating a nucleophilic residue for covalent catalysis. The residues form a charge-relay network to polarise and activate the nucleophile, which attacks the substrate, forming a covalent intermediate which is then hydrolysed to release the product and regenerate free enzyme. The nucleophile is most commonly a serine or cysteine amino acid, but occasionally threonine or even selenocysteine. The 3D structure of the enzyme brings together the triad residues in a precise orientation, even though they may be far apart in the sequence.

Chlorophyll b is a form of chlorophyll. Chlorophyll b helps in photosynthesis by absorbing light energy. It is more soluble than chlorophyll a in polar solvents because of its carbonyl group. Its color is green, and it primarily absorbs blue light.
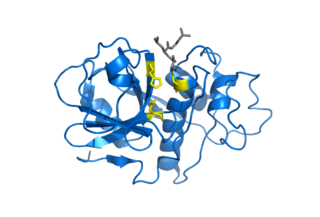
Cysteine proteases, also known as thiol proteases, are hydrolase enzymes that degrade proteins. These proteases share a common catalytic mechanism that involves a nucleophilic cysteine thiol in a catalytic triad or dyad.
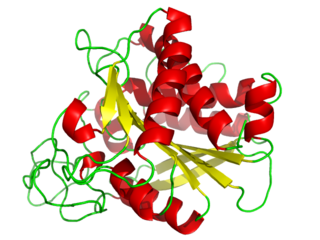
A carboxypeptidase is a protease enzyme that hydrolyzes (cleaves) a peptide bond at the carboxy-terminal (C-terminal) end of a protein or peptide. This is in contrast to an aminopeptidases, which cleave peptide bonds at the N-terminus of proteins. Humans, animals, bacteria and plants contain several types of carboxypeptidases that have diverse functions ranging from catabolism to protein maturation. At least two mechanisms have been discussed.
Serine hydrolases are one of the largest known enzyme classes comprising approximately ~200 enzymes or 1% of the genes in the human proteome. A defining characteristic of these enzymes is the presence of a particular serine at the active site, which is used for the hydrolysis of substrates. The hydrolysis of the ester or peptide bond proceeds in two steps. First, the acyl part of the substrate is transferred to the serine, making a new ester or amide bond and releasing the other part of the substrate is released. Later, in a slower step, the bond between the serine and the acyl group is hydrolyzed by water or hydroxide ion, regenerating free enzyme. Unlike other, non-catalytic, serines, the reactive serine of these hydrolases is typically activated by a proton relay involving a catalytic triad consisting of the serine, an acidic residue and a basic residue, although variations on this mechanism exist.
Branched-chain amino acid aminotransferase (BCAT), also known as branched-chain amino acid transaminase, is an aminotransferase enzyme (EC 2.6.1.42) which acts upon branched-chain amino acids (BCAAs). It is encoded by the BCAT2 gene in humans. The BCAT enzyme catalyzes the conversion of BCAAs and α-ketoglutarate into branched chain α-keto acids and glutamate.

The enzyme cutinase is a member of the hydrolase family. It catalyzes the following reaction:
The enzyme tannase (EC 3.1.1.20) catalyzes the following reaction:

In enzymology, an amidase (EC 3.5.1.4, acylamidase, acylase (misleading), amidohydrolase (ambiguous), deaminase (ambiguous), fatty acylamidase, N-acetylaminohydrolase (ambiguous)) is an enzyme that catalyzes the hydrolysis of an amide. In this way, the two substrates of this enzyme are an amide and H2O, whereas its two products are monocarboxylate and NH3.
In enzymology, an aminoacylase (EC 3.5.1.14) is an enzyme that catalyzes the chemical reaction

Protochlorophyllide, or monovinyl protochlorophyllide, is an intermediate in the biosynthesis of chlorophyll a. It lacks the phytol side-chain of chlorophyll and the reduced pyrrole in ring D. Protochlorophyllide is highly fluorescent; mutants that accumulate it glow red if irradiated with blue light. In angiosperms, the later steps which convert protochlorophyllide to chlorophyll are light-dependent, and such plants are pale (chlorotic) if grown in the darkness. Gymnosperms, algae, and photosynthetic bacteria have another, light-independent enzyme and grow green in the darkness as well.
In molecular biology, the red chlorophyll catabolite reductase family of proteins consists of several red chlorophyll catabolite reductase proteins. Red chlorophyll catabolite (RCC) reductase (RCCR) and pheophorbide (Pheide) a oxygenase (PaO) catalyse the key reaction of chlorophyll catabolism, porphyrin macrocycle cleavage of Pheide a to a primary fluorescent catabolite (pFCC).
Pheophorbide a oxygenase (EC 1.14.15.17, pheide a monooxygenase, pheide a oxygenase, PAO) is an enzyme with systematic name pheophorbide-a,NADPH:oxygen oxidoreductase (biladiene-forming). This enzyme catalyses the following chemical reaction

Chlorophyllide a and Chlorophyllide b are the biosynthetic precursors of chlorophyll a and chlorophyll b respectively. Their propionic acid groups are converted to phytyl esters by the enzyme chlorophyll synthase in the final step of the pathway. Thus the main interest in these chemical compounds has been in the study of chlorophyll biosynthesis in plants, algae and cyanobacteria. Chlorophyllide a is also an intermediate in the biosynthesis of bacteriochlorophylls.
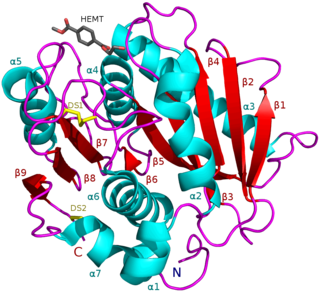
PETases are an esterase class of enzymes that catalyze the breakdown (via hydrolysis) of polyethylene terephthalate (PET) plastic to monomeric mono-2-hydroxyethyl terephthalate (MHET). The idealized chemical reaction is:















