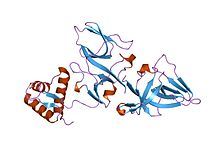
Mutagenesis is a process by which the genetic information of an organism is changed by the production of a mutation. It may occur spontaneously in nature, or as a result of exposure to mutagens. It can also be achieved experimentally using laboratory procedures. A mutagen is a mutation-causing agent, be it chemical or physical, which results in an increased rate of mutations in an organism's genetic code. In nature mutagenesis can lead to cancer and various heritable diseases, and it is also a driving force of evolution. Mutagenesis as a science was developed based on work done by Hermann Muller, Charlotte Auerbach and J. M. Robson in the first half of the 20th century.

A prophage is a bacteriophage genome that is integrated into the circular bacterial chromosome or exists as an extrachromosomal plasmid within the bacterial cell. Integration of prophages into the bacterial host is the characteristic step of the lysogenic cycle of temperate phages. Prophages remain latent in the genome through multiple cell divisions until activation by an external factor, such as UV light, leading to production of new phage particles that will lyse the cell and spread. As ubiquitous mobile genetic elements, prophages play important roles in bacterial genetics and evolution, such as in the acquisition of virulence factors.

The SOS response is a global response to DNA damage in which the cell cycle is arrested and DNA repair and mutagenesis are induced. The system involves the RecA protein. The RecA protein, stimulated by single-stranded DNA, is involved in the inactivation of the repressor (LexA) of SOS response genes thereby inducing the response. It is an error-prone repair system that contributes significantly to DNA changes observed in a wide range of species.
Integrons are genetic mechanisms that allow bacteria to adapt and evolve rapidly through the stockpiling and expression of new genes. These genes are embedded in a specific genetic structure called gene cassette that generally carries one promoterless open reading frame (ORF) together with a recombination site (attC). Integron cassettes are incorporated to the attI site of the integron platform by site-specific recombination reactions mediated by the integrase.
DNA gyrase, or simply gyrase, is an enzyme within the class of topoisomerase and is a subclass of Type II topoisomerases that reduces topological strain in an ATP dependent manner while double-stranded DNA is being unwound by elongating RNA-polymerase or by helicase in front of the progressing replication fork. It is the only known enzyme to actively contribute negative supercoiling to DNA, while it also is capable of relaxing positive supercoils. It does so by looping the template to form a crossing, then cutting one of the double helices and passing the other through it before releasing the break, changing the linking number by two in each enzymatic step. This process occurs in bacteria, whose single circular DNA is cut by DNA gyrase and the two ends are then twisted around each other to form supercoils. Gyrase is also found in eukaryotic plastids: it has been found in the apicoplast of the malarial parasite Plasmodium falciparum and in chloroplasts of several plants. Bacterial DNA gyrase is the target of many antibiotics, including nalidixic acid, novobiocin, albicidin, and ciprofloxacin.

RecA is a 38 kilodalton protein essential for the repair and maintenance of DNA. A RecA structural and functional homolog has been found in every species in which one has been seriously sought and serves as an archetype for this class of homologous DNA repair proteins. The homologous protein is called RAD51 in eukaryotes and RadA in archaea.
A DNA-binding domain (DBD) is an independently folded protein domain that contains at least one structural motif that recognizes double- or single-stranded DNA. A DBD can recognize a specific DNA sequence or have a general affinity to DNA. Some DNA-binding domains may also include nucleic acids in their folded structure.
SOS box is the operator to which the LexA repressor binds to repress the transcription of SOS-induced proteins. SOS boxes are found near the promoter of various genes.

Tet Repressor proteins are proteins playing an important role in conferring antibiotic resistance to large categories of bacterial species.
The rpoB gene encodes the β subunit of bacterial RNA polymerase and the homologous plastid-encoded RNA polymerase (PEP). It codes for 1342 amino acids in E. coli, making it the second-largest polypeptide in the bacterial cell. It is targeted by the rifamycin family of antibacterials, such as rifampin. Mutations in rpoB that confer resistance to rifamycins do so by altering the protein's drug-binding residues, thereby reducing affinity for these antibiotics.
Topoisomerase inhibitors are chemical compounds that block the action of topoisomerases, which are broken into two broad subtypes: type I topoisomerases (TopI) and type II topoisomerases (TopII). Topoisomerase plays important roles in cellular reproduction and DNA organization, as they mediate the cleavage of single and double stranded DNA to relax supercoils, untangle catenanes, and condense chromosomes in eukaryotic cells. Topoisomerase inhibitors influence these essential cellular processes. Some topoisomerase inhibitors prevent topoisomerases from performing DNA strand breaks while others, deemed topoisomerase poisons, associate with topoisomerase-DNA complexes and prevent the re-ligation step of the topoisomerase mechanism. These topoisomerase-DNA-inhibitor complexes are cytotoxic agents, as the un-repaired single- and double stranded DNA breaks they cause can lead to apoptosis and cell death. Because of this ability to induce apoptosis, topoisomerase inhibitors have gained interest as therapeutics against infectious and cancerous cells.

Type II topoisomerases are topoisomerases that cut both strands of the DNA helix simultaneously in order to manage DNA tangles and supercoils. They use the hydrolysis of ATP, unlike Type I topoisomerase. In this process, these enzymes change the linking number of circular DNA by ±2. Topoisomerases are ubiquitous enzymes, found in all living organisms.

Aminocoumarin is a class of antibiotics that act by an inhibition of the DNA gyrase enzyme involved in the cell division in bacteria. They are derived from Streptomyces species, whose best-known representative – Streptomyces coelicolor – was completely sequenced in 2002. The aminocoumarin antibiotics include:
SmeT is a transcriptional repressor protein of 24.6 kDa, found in the pathogenic bacteria Stenotrophomonas maltophilia. SmeT is responsible for the regulation of the Multidrug Resistance (MDR) efflux pump, SmeDEF, that gives the bacteria resistance to several antibiotics including macrolides, TMP/SMX, tetracycline, chloramphenicol, quinolones and erythromycin. SmeT is encoded 223 bp upstream of SmeDEF, with just 56 base pairs between their transcription start sites and an overlapping region between the promoters. The production of the SmeT protein downregulates its own transcription, along with that of the efflux pump by sterically hindering the binding of RNA Polymerase to the DNA. SmeDEF was the first MDR pump discovered in the S. maltophilia species. The pump is named by its different parts: SmeE, the transporter itself that spans the plasma membrane, SmeF, the protein on the outer portion of the membrane, and SmeD, a membrane fusion protein. On general purpose media and no selectors, the genes for MDR pumps are typically not expressed, and the repressor is found bound to the DNA. In fact, mutations in SmeT that lead to overexpression of SmeDEF can pose fitness challenges to the bacteria. However, this overexpression has been identified in the bacterium and may pose a threat to our health.
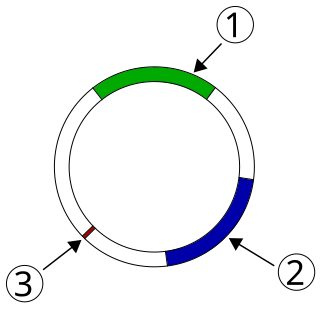
Plasmid-mediated resistance is the transfer of antibiotic resistance genes which are carried on plasmids. Plasmids possess mechanisms that ensure their independent replication as well as those that regulate their replication number and guarantee stable inheritance during cell division. By the conjugation process, they can stimulate lateral transfer between bacteria from various genera and kingdoms. Numerous plasmids contain addiction-inducing systems that are typically based on toxin-antitoxin factors and capable of killing daughter cells that don't inherit the plasmid during cell division. Plasmids often carry multiple antibiotic resistance genes, contributing to the spread of multidrug-resistance (MDR). Antibiotic resistance mediated by MDR plasmids severely limits the treatment options for the infections caused by Gram-negative bacteria, especially family Enterobacteriaceae. The global spread of MDR plasmids has been enhanced by selective pressure from antimicrobial medications used in medical facilities and when raising animals for food.

Quinolone antibiotics constitute a large group of broad-spectrum bacteriocidals that share a bicyclic core structure related to the substance 4-quinolone. They are used in human and veterinary medicine to treat bacterial infections, as well as in animal husbandry, specifically poultry production.
Bacterial morphological plasticity refers to changes in the shape and size that bacterial cells undergo when they encounter stressful environments. Although bacteria have evolved complex molecular strategies to maintain their shape, many are able to alter their shape as a survival strategy in response to protist predators, antibiotics, the immune response, and other threats.

Neisseria gonorrhoeae, the bacterium that causes the sexually transmitted infection gonorrhea, has developed antibiotic resistance to many antibiotics. The bacteria was first identified in 1879.
DNA Polymerase V is a polymerase enzyme involved in DNA repair mechanisms in bacteria, such as Escherichia coli. It is composed of a UmuD' homodimer and a UmuC monomer, forming the UmuD'2C protein complex. It is part of the Y-family of DNA Polymerases, which are capable of performing DNA translesion synthesis (TLS). Translesion polymerases bypass DNA damage lesions during DNA replication - if a lesion is not repaired or bypassed the replication fork can stall and lead to cell death. However, Y polymerases have low sequence fidelity during replication. When the UmuC and UmuD' proteins were initially discovered in E. coli, they were thought to be agents that inhibit faithful DNA replication and caused DNA synthesis to have high mutation rates after exposure to UV-light. The polymerase function of Pol V was not discovered until the late 1990s when UmuC was successfully extracted, consequent experiments unequivocally proved UmuD'2C is a polymerase. This finding lead to the detection of many Pol V orthologs and the discovery of the Y-family of polymerases.

Ivan Erill is a Spanish computational biologist known for his research in comparative genomics and molecular microbiology. His work focuses primarily on bacterial comparative genomics, through the development of computational methods for analyzing regulatory networks and their evolution.