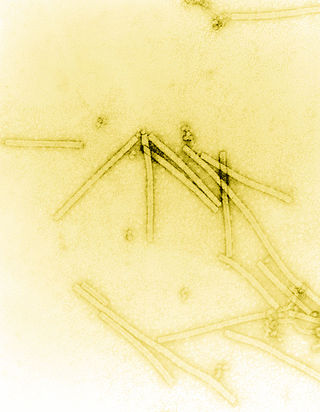
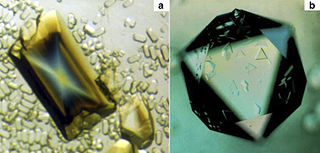
Tobacco mosaic virus (TMV) is a positive-sense single-stranded RNA virus species in the genus Tobamovirus that infects a wide range of plants, especially tobacco and other members of the family Solanaceae. The infection causes characteristic patterns, such as "mosaic"-like mottling and discoloration on the leaves. TMV was the first virus to be discovered. Although it was known from the late 19th century that a non-bacterial infectious disease was damaging tobacco crops, it was not until 1930 that the infectious agent was determined to be a virus. It is the first pathogen identified as a virus. The virus was crystallised by Wendell Meredith Stanley. It has a similar size to the largest synthetic molecule, known as PG5.
A satellite is a subviral agent that depends on the coinfection of a host cell with a helper virus for its replication. Satellites can be divided into two major classes: satellite viruses and satellite nucleic acids. Satellite viruses, which are most commonly associated with plants, are also found in mammals, arthropods, and bacteria. They encode structural proteins to enclose their genetic material, which are therefore distinct from the structural proteins of their helper viruses. Satellite nucleic acids, in contrast, do not encode their own structural proteins, but instead are encapsulated by proteins encoded by their helper viruses. The genomes of satellites range upward from 359 nucleotides in length for satellite tobacco ringspot virus RNA (STobRV).

Geminiviridae is a family of plant viruses that encode their genetic information on a circular genome of single-stranded (ss) DNA. There are 520 species in this family, assigned to 14 genera. Diseases associated with this family include: bright yellow mosaic, yellow mosaic, yellow mottle, leaf curling, stunting, streaks, reduced yields. They have single-stranded circular DNA genomes encoding genes that diverge in both directions from a virion strand origin of replication. According to the Baltimore classification they are considered class II viruses. It is the largest known family of single stranded DNA viruses.

Tombusviridae is a family of single-stranded positive sense RNA plant viruses. There are three subfamilies, 17 genera, and 95 species in this family. The name is derived from Tomato bushy stunt virus (TBSV).

Tomato bushy stunt virus (TBSV) is a virus of the tombusvirus family. It was first reported in tomatoes in 1935 and primarily affects vegetable crops, though it is not generally considered an economically significant plant pathogen. Depending upon the host, TBSV causes stunting of growth, leaf mottling, and deformed or absent fruit. The virus is likely to be soil-borne in the natural setting, but can also be transmitted mechanically, for example through contaminated cutting tools. TBSV has been used as a model system in virology research on the life cycle of plant viruses, particularly in experimental infections of the model host plant Nicotiana benthamiana.

Potyvirus is a genus of positive-strand RNA viruses in the family Potyviridae. Plants serve as natural hosts. Like begomoviruses, members of this genus may cause significant losses in agricultural, pastoral, horticultural, and ornamental crops. More than 200 species of aphids spread potyviruses, and most are from the subfamily Aphidinae. The genus contains 190 species and potyviruses account for about thirty percent of all currently known plant viruses.
Tombusvirus is a genus of viruses, in the family Tombusviridae. Plants serve as natural hosts. There are 17 species in this genus. Symptoms associated with this genus include mosaic. The name of the genus comes from Tomato bushy stunt virus.
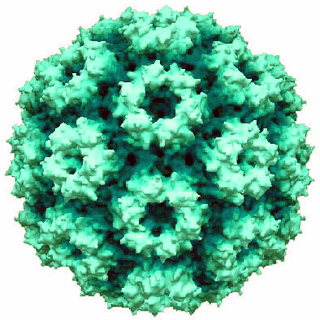
Cowpea chlorotic mottle virus, known by the abbreviation CCMV, is a virus that specifically infects the cowpea plant, or black-eyed pea. The leaves of infected plants develop yellow spots, hence the name "chlorotic". Similar to its "brother" virus, Cowpea mosaic virus (CPMV), CCMV is produced in high yield in plants. In the natural host, viral particles can be produced at 1–2 mg per gram of infected leaf tissue. Belonging to the bromovirus genus, cowpea chlorotic mottle virus (CCMV) is a small spherical plant virus. Other members of this genus include the brome mosaic virus (BMV) and the broad bean mottle virus (BBMV).
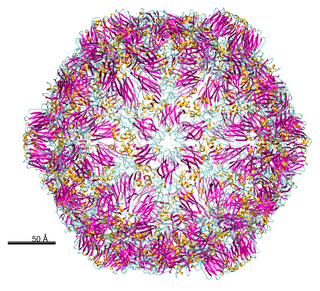
Nepovirus is a genus of viruses in the order Picornavirales, in the family Secoviridae, in the subfamily Comovirinae. Plants serve as natural hosts. There are 40 species in this genus. Nepoviruses, unlike the other two genera in the subfamily Comovirinae, are transmitted by nematodes.

Ilarvirus is a genus of positive-strand RNA viruses in the family Bromoviridae. Plants serve as natural hosts. There are 22 species in this genus.

Sobemovirus is a genus of non-enveloped, positive-strand RNA viruses which infect plants. Plants serve as natural hosts. There are 21 species in this genus. Diseases associated with this genus include: mosaics and mottles.

Potexvirus is a genus of pathogenic viruses in the order Tymovirales, in the family Alphaflexiviridae. Plants serve as natural hosts. There are 48 species in this genus, three of which are assigned to a subgenus. Diseases associated with this genus include: mosaic and ringspot symptoms. The genus name comes from POTato virus X).

Carlavirus, formerly known as the "Carnation latent virus group", is a genus of viruses in the order Tymovirales, in the family Betaflexiviridae. Plants serve as natural hosts. There are 53 species in this genus. Diseases associated with this genus include: mosaic and ringspot symptoms.

Cucumovirus is a genus of viruses, in the family Bromoviridae. Plants serve as natural hosts. There are four species in this genus.

Virgaviridae is a family of positive-strand RNA viruses. Plants serve as natural hosts. The name of the family is derived from the Latin word virga (rod), as all viruses in this family are rod-shaped. There are currently 59 species in this family, divided among seven genera.
Ipomovirus is a genus of positive-strand RNA viruses in the family Potyviridae. Member viruses infect plants and are transmitted by whiteflies. The name of the genus is derived from Ipomoea – the generic name of sweet potato. There are seven species in this genus.

Bromovirus is a genus of viruses, in the family Bromoviridae. Plants serve as natural hosts. There are six species in this genus.

Caulimovirus is a genus of viruses, in the family Caulimoviridae order Ortervirales. They are para-retroviruses with dsDNA and plants as their host. There are 12 species in this genus. Diseases associated with this genus include: vein-clearing or banding mosaic.

Tymovirus is a genus of viruses in the order Tymovirales, in the family Tymoviridae. Plants serve as natural hosts. There are 28 species in this genus.
Riboviria is a realm of viruses that includes all viruses that use a homologous RNA-dependent polymerase for replication. It includes RNA viruses that encode an RNA-dependent RNA polymerase, as well as reverse-transcribing viruses that encode an RNA-dependent DNA polymerase. RNA-dependent RNA polymerase (RdRp), also called RNA replicase, produces RNA from RNA. RNA-dependent DNA polymerase (RdDp), also called reverse transcriptase (RT), produces DNA from RNA. These enzymes are essential for replicating the viral genome and transcribing viral genes into messenger RNA (mRNA) for translation of viral proteins.



















