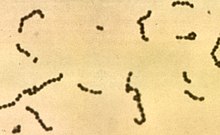Group A streptococcal infections are a number of infections with Streptococcus pyogenes, a group A streptococcus (GAS). S. pyogenes is a species of beta-hemolytic Gram-positive bacteria that is responsible for a wide range of infections that are mostly common and fairly mild. If the bacteria enters the bloodstream, the infection can become severe and life-threatening, and is called an invasive GAS (iGAS).

Streptococcus pyogenes is a species of Gram-positive, aerotolerant bacteria in the genus Streptococcus. These bacteria are extracellular, and made up of non-motile and non-sporing cocci that tend to link in chains. They are clinically important for humans, as they are an infrequent, but usually pathogenic, part of the skin microbiota that can cause group A streptococcal infection. S. pyogenes is the predominant species harboring the Lancefield group A antigen, and is often called group A Streptococcus (GAS). However, both Streptococcus dysgalactiae and the Streptococcus anginosus group can possess group A antigen as well. Group A streptococci, when grown on blood agar, typically produce small (2–3 mm) zones of beta-hemolysis, a complete destruction of red blood cells. The name group A (beta-hemolytic) Streptococcus is thus also used.
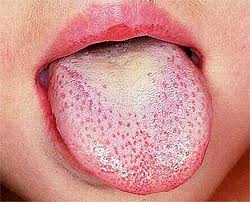
Scarlet fever, also known as scarlatina, is an infectious disease caused by Streptococcus pyogenes, a Group A streptococcus (GAS). It most commonly affects children between five and 15 years of age. The signs and symptoms include a sore throat, fever, headache, swollen lymph nodes, and a characteristic rash. The face is flushed and the rash is red and blanching. It typically feels like sandpaper and the tongue may be red and bumpy. The rash occurs as a result of capillary damage by exotoxins produced by S.pyogenes. On darker-pigmented skin the rash may be hard to discern.

The viridans streptococci are a large group of commensal streptococcal Gram-positive bacteria species that are α-hemolytic, producing a green coloration on blood agar plates, although some species in this group are actually γ-hemolytic, meaning they produce no change on blood agar. The pseudo-taxonomic term "Streptococcus viridans" is often used to refer to this group of species, but writers who do not like to use the pseudotaxonomic term prefer the terms viridans streptococci, viridans group streptococci (VGS), or viridans streptococcal species.

Hemolysis is the breakdown of red blood cells. The ability of bacterial colonies to induce hemolysis when grown on blood agar is used to classify certain microorganisms. This is particularly useful in classifying streptococcal species. A substance that causes hemolysis is called a hemolysin.

Rebecca Craighill Lancefield was a prominent American microbiologist. She joined the Rockefeller Institute for Medical Research in New York in 1918, and was associated with that institute throughout her long and outstanding career. Her bibliography comprises more than 50 publications published over 60 years.

Arcanobacterium haemolyticum is a species of bacteria classified as a gram-positive bacillus. It is catalase-negative, facultative anaerobic, beta-hemolytic, and not motile. It has been known to cause head and neck infections, pharyngitis, and sinusitis.
Anti-streptolysin O is the antibody made against streptolysin O, an immunogenic, oxygen-labile streptococcal hemolytic exotoxin produced by most strains of group A and many strains of groups C and G Streptococcus bacteria. The "O" in the name stands for oxygen-labile; the other related toxin being oxygen-stable streptolysin-S. The main function of streptolysin O is to cause hemolysis —in particular, beta-hemolysis.
Streptolysins are two homogenous exotoxins from Streptococcus pyogenes. Types include streptolysin O, which is oxygen-labile, and streptolysin S, which is oxygen-stable.
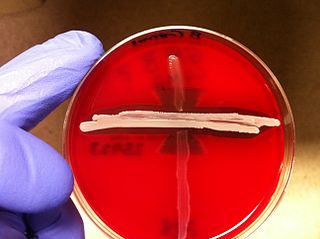
The CAMP test (Christie–Atkins–Munch-Petersen) is a test to identify group B β-hemolytic streptococci based on their formation of a substance, CAMP factor, that enlarges the area of hemolysis formed by the β-hemolysin elaborated from Staphylococcus aureus.

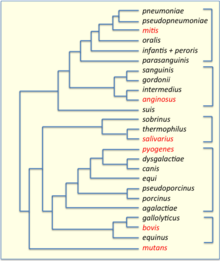
Streptococcus anginosus is a species of Streptococcus. This species, Streptococcus intermedius, and Streptococcus constellatus constitute the anginosus group, which is sometimes also referred to as the milleri group after the previously assumed but later refuted idea of a single species Streptococcus milleri. Phylogenetic relatedness of S. anginosus, S. constellatus, and S. intermedius has been confirmed by rRNA sequence analysis.

Streptococcus dysgalactiae is a gram positive, beta-haemolytic, coccal bacterium belonging to the family Streptococcaceae. It is capable of infecting both humans and animals, but is most frequently encountered as a commensal of the alimentary tract, genital tract, or less commonly, as a part of the skin flora. The clinical manifestations in human disease range from superficial skin-infections and tonsillitis, to severe necrotising fasciitis and bacteraemia. The incidence of invasive disease has been reported to be rising. Several different animal species are susceptible to infection by S. dysgalactiae, but bovine mastitis and infectious arthritis in lambs have been most frequently reported.

Streptococcal intertrigo is a skin condition that is secondary to a streptococcal bacterial infection. It is often seen in infants and young children and can be characterized by a fiery-red color of the skin, foul odor with an absence of satellite lesions, and skin softening in the neck, armpits or folds of the groin. Newborn children and infants commonly develop intertrigo because of physical features such as deep skin folds, short neck, and flexed posture. Prompt diagnosis by a medical professional and treatment with topical and/or oral antibiotics can effectively relieve symptoms.

Streptococcus iniae is a species of Gram-positive, sphere-shaped bacterium belonging to the genus Streptococcus. Since its isolation from an Amazon freshwater dolphin in the 1970s, S. iniae has emerged as a leading fish pathogen in aquaculture operations worldwide, resulting in over US$100M in annual losses. Since its discovery, S. iniae infections have been reported in at least 27 species of cultured or wild fish from around the world. Freshwater and saltwater fish including tilapia, red drum, hybrid striped bass, and rainbow trout are among those susceptible to infection by S. iniae. Infections in fish manifest as meningoencephalitis, skin lesions, and septicemia.
Perianal cellulitis, also known as perianitis or perianal streptococcal dermatitis, is a bacterial infection affecting the lower layers of the skin (cellulitis) around the anus. It presents as bright redness in the skin and can be accompanied by pain, difficulty defecating, itching, and bleeding. This disease is considered a complicated skin and soft tissue infection (cSSTI) because of the involvement of the deeper soft tissues.
Bacteriophage T12 is a bacteriophage that infects Streptococcus pyogenes bacteria. It is a proposed species of the family Siphoviridae in the order Caudovirales also known as tailed viruses. It converts a harmless strain of bacteria into a virulent strain. It carries the speA gene which codes for erythrogenic toxin A. speA is also known as streptococcal pyogenic exotoxin A, scarlet fever toxin A, or even scarlatinal toxin. Note that the name of the gene "speA" is italicized; the name of the toxin "speA" is not italicized. Erythrogenic toxin A converts a harmless, non-virulent strain of Streptococcus pyogenes to a virulent strain through lysogeny, a life cycle which is characterized by the ability of the genome to become a part of the host cell and be stably maintained there for generations. Phages with a lysogenic life cycle are also called temperate phages. Bacteriophage T12, proposed member of family Siphoviridae including related speA-carrying bacteriophages, is also a prototypic phage for all the speA-carrying phages of Streptococcus pyogenes, meaning that its genome is the prototype for the genomes of all such phages of S. pyogenes. It is the main suspect as the cause of scarlet fever, an infectious disease that affects small children.

Lancefield grouping is a system of classification that classifies catalase-negative Gram-positive cocci based on the carbohydrate composition of bacterial antigens found on their cell walls. The system, created by Rebecca Lancefield, was historically used to organize the various members of the family Streptococcaceae, which includes the genera Lactococcus and Streptococcus, but now is largely superfluous due to explosive growth in the number of streptococcal species identified since the 1970s. However, it has retained some clinical usefulness even after the taxonomic changes, and as of 2018, Lancefield designations are still often used to communicate medical microbiological test results.

In microbiology, colonial morphology refers to the visual appearance of bacterial or fungal colonies on an agar plate. Examining colonial morphology is the first step in the identification of an unknown microbe. The systematic assessment of the colonies' appearance, focusing on aspects like size, shape, colour, opacity, and consistency, provides clues to the identity of the organism, allowing microbiologists to select appropriate tests to provide a definitive identification.
Columbia Nalidixic Acid (CNA) agar is a growth medium used for the isolation and cultivation of bacteria from clinical and non-clinical specimens. CNA agar contains antibiotics that inhibit Gram-negative organisms, aiding in the selective isolation of Gram-positive bacteria. Gram-positive organisms that grow on the media can be differentiated on the basis of hemolysis.
Streptococcosis is an infectious disease caused by bacteria of the genus Steptococcus. This disease is most common among horses, guinea pigs, dogs, cats, and fish with symptoms varying based on the streptococcal species involved. In humans, this disease typically involves a throat infection and is called streptococcal pharyngitis or strep throat.