Related Research Articles

A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" base pairs allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins can recognize specific base-pairing patterns that identify particular regulatory regions of genes.

Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver.

Protein biosynthesis is a core biological process, occurring inside cells, balancing the loss of cellular proteins through the production of new proteins. Proteins perform a number of critical functions as enzymes, structural proteins or hormones. Protein synthesis is a very similar process for both prokaryotes and eukaryotes but there are some distinct differences.
Deamination is the removal of an amino group from a molecule. Enzymes that catalyse this reaction are called deaminases.
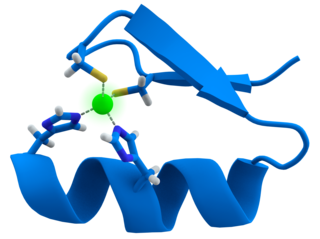
A zinc finger is a small protein structural motif that is characterized by the coordination of one or more zinc ions (Zn2+) in order to stabilize the fold. It was originally coined to describe the finger-like appearance of a hypothesized structure from the African clawed frog (Xenopus laevis) transcription factor IIIA. However, it has been found to encompass a wide variety of differing protein structures in eukaryotic cells. Xenopus laevis TFIIIA was originally demonstrated to contain zinc and require the metal for function in 1983, the first such reported zinc requirement for a gene regulatory protein followed soon thereafter by the Krüppel factor in Drosophila. It often appears as a metal-binding domain in multi-domain proteins.
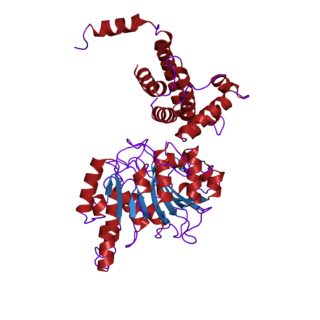
In biochemistry, the DNA methyltransferase family of enzymes catalyze the transfer of a methyl group to DNA. DNA methylation serves a wide variety of biological functions. All the known DNA methyltransferases use S-adenosyl methionine (SAM) as the methyl donor.

EGR-1 also known as ZNF268 or NGFI-A is a protein that in humans is encoded by the EGR1 gene.
The C7.GAT protein is a zinc finger protein based on the C7 protein. It features an alternative zinc finger 3 alpha helix sequence, preventing the target site overlap caused by the aspartic acid residue of the finger 3 of C7. The sequence of this third finger is TSG-N-LVR according to the single letter amino acid code. As the name suggest, the target site of finger 3 is altered to 5'-GAT-3', giving the overall protein a target of 5'-GCGTGGGAT-3'.
Therapeutic gene modulation refers to the practice of altering the expression of a gene at one of various stages, with a view to alleviate some form of ailment. It differs from gene therapy in that gene modulation seeks to alter the expression of an endogenous gene whereas gene therapy concerns the introduction of a gene whose product aids the recipient directly.
In a zinc finger protein, certain sequences of amino acid residues are able to recognise and bind to an extended target-site of four or even five nucleotides When this occurs in a ZFP in which the three-nucleotide subsites are contiguous, one zinc finger interferes with the target-site of the zinc finger adjacent to it, a situation known as target-site overlap. For example, a zinc finger containing arginine at position -1 and aspartic acid at position 2 along its alpha-helix will recognise an extended sequence of four nucleotides of the sequence 5'-NNG(G/T)-3'. The hydrogen bond between Asp2 and the N4 of either a cytosine or adenine base paired to the guanine or thymine, respectively defines these two nucleotides at the 3' position, defining a sequence that overlaps into the subsite of any zinc finger that may be attached N-terminally.
Protein metabolism denotes the various biochemical processes responsible for the synthesis of proteins and amino acids (anabolism), and the breakdown of proteins by catabolism.

PDE3 is a phosphodiesterase. The PDEs belong to at least eleven related gene families, which are different in their primary structure, substrate affinity, responses to effectors, and regulation mechanism. Most of the PDE families are composed of more than one gene. PDE3 is clinically significant because of its role in regulating heart muscle, vascular smooth muscle and platelet aggregation. PDE3 inhibitors have been developed as pharmaceuticals, but their use is limited by arrhythmic effects and they can increase mortality in some applications.

The PDE2 enzyme is one of 21 different phosphodiesterases (PDE) found in mammals. These different PDEs can be subdivided to 11 families. The different PDEs of the same family are functionally related despite the fact that their amino acid sequences show considerable divergence. The PDEs have different substrate specificities. Some are cAMP selective hydrolases, others are cGMP selective hydrolases and the rest can hydrolyse both cAMP and cGMP.
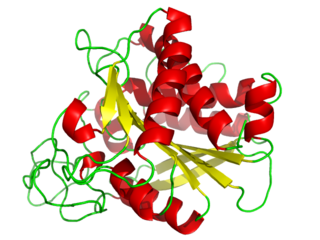
Carboxypeptidase A usually refers to the pancreatic exopeptidase that hydrolyzes peptide bonds of C-terminal residues with aromatic or aliphatic side-chains. Most scientists in the field now refer to this enzyme as CPA1, and to a related pancreatic carboxypeptidase as CPA2.
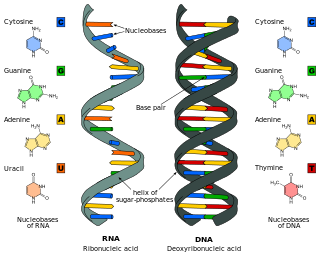
Nucleic acid analogues are compounds which are analogous to naturally occurring RNA and DNA, used in medicine and in molecular biology research. Nucleic acids are chains of nucleotides, which are composed of three parts: a phosphate backbone, a pentose sugar, either ribose or deoxyribose, and one of four nucleobases. An analogue may have any of these altered. Typically the analogue nucleobases confer, among other things, different base pairing and base stacking properties. Examples include universal bases, which can pair with all four canonical bases, and phosphate-sugar backbone analogues such as PNA, which affect the properties of the chain . Nucleic acid analogues are also called Xeno Nucleic Acid and represent one of the main pillars of xenobiology, the design of new-to-nature forms of life based on alternative biochemistries.
Zinc finger protein chimera are chimeric proteins composed of a DNA-binding zinc finger protein domain and another domain through which the protein exerts its effect. The effector domain may be a transcriptional activator (A) or repressor (R), a methylation domain (M) or a nuclease (N).

Zinc finger protein 804A is a protein that in humans is encoded by the ZNF804A gene. The human gene maps to chromosome 2 q32.1 and consists of 4 exons that code for a protein of 1210 amino acids.

Nucleic acid structure refers to the structure of nucleic acids such as DNA and RNA. Chemically speaking, DNA and RNA are very similar. Nucleic acid structure is often divided into four different levels: primary, secondary, tertiary, and quaternary.

The WRKY domain is found in the WRKY transcription factor family, a class of transcription factors. The WRKY domain is found almost exclusively in plants although WRKY genes appear present in some diplomonads, social amoebae and other amoebozoa, and fungi incertae sedis. They appear absent in other non-plant species. WRKY transcription factors have been a significant area of plant research for the past 20 years. The WRKY DNA-binding domain recognizes the W-box (T)TGAC(C/T) cis-regulatory element.
Since antiretroviral therapy requires a lifelong treatment regimen, research to find more permanent cures for HIV infection is currently underway. It is possible to synthesize zinc finger nucleotides with zinc finger components that selectively bind to specific portions of DNA. Conceptually, targeting and editing could focus on host cellular co-receptors for HIV or on proviral HIV DNA.
References
- Wu H, Yang WP, Barbas CF (January 1995). "Building zinc fingers by selection: toward a therapeutic application". Proc. Natl. Acad. Sci. U.S.A. 92 (2): 344–8. doi: 10.1073/pnas.92.2.344 . PMC 42736 . PMID 7831288.