Function
The protein encoded by this gene belongs to the EGR family of Cys2His2-type zinc finger proteins. It is a nuclear protein and functions as a transcriptional regulator. The products of target genes it activates are required for differentiation and mitogenesis. Studies suggest this is a tumor suppressor gene. [5]
It has a distinct pattern of expression in the brain, and its induction has been shown to be associated with neuronal activity. Several studies suggest it has a role in neuronal plasticity. [6]
EGR-1 is an important transcription factor in memory formation. It has an essential role in brain neuron epigenetic reprogramming. EGR-1 recruits the TET1 protein that initiates a pathway of DNA demethylation. [7] Removing DNA methylation marks allows the activation of downstream genes. EGR-1, together with TET1, is employed in programming the distribution of methylation sites on brain DNA during brain development, in learning and in long-term neuronal plasticity. EGR-1 has also been found to regulate the expression of VAMP2 (a protein important for synaptic exocytosis). [8]
Beside its function in the nervous system, there is significant evidence that EGR-1 along with its paralog EGR-2 is induced in fibrotic diseases has key functions in fibrinogenesis and is necessary for experimentally induced fibrosis in mice. [9]
It may also be involved in ovarian function [10]
Structure
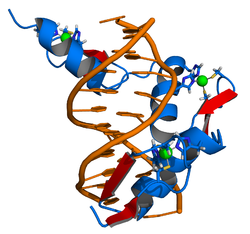
The DNA-binding domain of EGR-1 consists of three zinc finger domains of the Cys2His2 type. The amino acid structure of the EGR-1 zinc finger domain is given in this table, using the single letter amino acid code. The fingers 1 to 3 are indicated by f1 - f3. The numbers are in reference to the residues (amino acids) of alpha helix (there is no zero). The residues marked 'x' are not part of the zinc fingers, but rather serve to connect them all together.
| | | | | | | | | | | | | | | | | | | | -1 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | | | x | x | x | x | x |
| f1 | M | A | E | E | R | P | Y | A | C | P | V | E | S | C | D | R | R | F | S | R | S | D | E | L | T | R | H | I | R | I | H | T | G | Q | K | P |
| f2 | | | | | | | F | Q | C | R | I | - | - | C | M | R | N | F | S | R | S | D | H | L | T | T | H | I | R | T | H | T | G | E | K | P |
| f3 | | | | | | | F | A | C | D | I | - | - | C | G | R | K | F | A | R | S | D | E | R | K | R | H | T | K | I | H | L | R | Q | K | D | |
Amino acid key: Alanine (Ala, A), Arginine (Arg, R), Asparagine (Asn, N), Aspartic acid (Asp, D), Cysteine (Cys, C), Glutamic acid (Glu, E), Glutamine (Gln, Q), Glycine (Gly, G), Histidine (His, H), Isoleucine (Ile, I), Leucine (Leu, L), Lysine (Lys, K), Methionine (Met, M), Phenylalanine (Phe, F), Proline (Pro, P), Serine (Ser, S), Threonine (Thr, T), Tryptophan (Trp, W), Tyrosine (Tyr, Y), Valine (Val, V)
The crystal structure of DNA bound by the zinc finger domain of EGR-1 was solved in 1991, which greatly aided early research in zinc finger DNA-binding domains. [11]
The human EGR-1 protein contains (in its unprocessed form) 543 amino acids with a molecular weight of 57.5 kDa, and the gene is located on the chromosome 5.
This page is based on this
Wikipedia article Text is available under the
CC BY-SA 4.0 license; additional terms may apply.
Images, videos and audio are available under their respective licenses.